Journal of
eISSN: 2373-4310


Case Report Volume 13 Issue 3
Department of Food Nutrition and Health, Central State University, USA
Correspondence: Pratibha Gupta, Ph.D., Research Associate Professor, Food, Nutrition and Health, Project Grant Director (funded by USDA/NIFA/CBG 1890), Core Director, Human Health Assessment and Nutrigenomics Lab, Agricultural Research Development Program A.R.D.P, College of Engineering, Science, Technology, and Agriculture (C.E.S.T.A.), Central State University, 1400 Brush Row Road, PO Box 100,4 Wilberforce, Ohio 45384 USA, 1890 LAND-GRANT INSTITUTION
Received: August 15, 2025 | Published: September 1, 2025
Citation: Gupta P, Charles S, Munroe S. Nutrition in action with nutrigenomics: genomic profiles for obesity risk reduction. J Nutr Health Food Eng. 2025;13(3):21-25 DOI: 10.15406/jnhfe.2025.13.00378
The integration of nutrigenomics into public health has the potential to revolutionize obesity prevention and treatment by providing personalized dietary recommendations tailored to an individual's unique genetic profile. By understanding how genetic variations affect responses to different nutrients, nutrigenomics enables targeted and effective weight management strategies. This approach has shown promise in reducing obesity risk and improving overall health outcomes. Our review highlights the current state of nutrigenomics in obesity risk reduction, including the key genes involved, benefits, and applications. We discuss the potential of nutrigenomics to optimize nutrient intake, prevent deficiencies, and promote overall health and wellbeing. Furthermore, we explore the challenges and limitations of implementing nutrigenomics in public health, including accessibility, affordability, and the need for further research. Ultimately, nutrigenomics has the potential to transform the way we approach nutrition and obesity prevention, providing a personalized and targeted approach to improving health outcomes.
Keywords: nutrigenomics, obesity, genomic profiles, personalized nutrition, weight management, public health
Obesity has emerged as one of the most critical global public health challenges of the 21st century. Defined by the World Health Organization (WHO) as an abnormal or excessive fat accumulation that presents a risk to health, obesity affects over two billion adults and more than 340 million children worldwide. This complex condition is a major risk factor for a multitude of chronic diseases, including type 2 diabetes, cardiovascular disease, non-alcoholic fatty liver disease, certain cancers, and musculoskeletal disorders. The socio-economic impact of obesity is immense, not only due to rising healthcare costs but also through reduced workforce productivity, increased disability, and social stigma. Despite widespread awareness campaigns and public health interventions emphasizing caloric balance through diet and exercise, obesity rates continue to rise. This suggests that conventional approaches to the prevention and treatment of obesity, which often adopt a "one-size-fits-all" model, are insufficient in addressing the underlying complexity of the condition. The traditional paradigm assumes that all individuals respond similarly to the same dietary and physical activity interventions, neglecting the biological variability that exists among individuals. Recent advances in genomic science have revolutionized our understanding of how genetic and environmental factors interact to influence metabolic health. In particular, the emerging field of nutrigenomics, which studies how nutrients and dietary patterns influence gene expression, offers a promising, personalized approach to combating obesity. Nutrigenomics bridges molecular biology, nutritional science, and genomics to examine how gene-diet interactions influence metabolic pathways, appetite regulation, lipid metabolism, insulin sensitivity, and inflammatory responses. Individuals possess unique genetic profiles characterized by single-nucleotide polymorphisms (SNPs) and other genetic variants that affect how their bodies metabolize food, respond to exercise, and store fat. By integrating genetic data with dietary habits, nutrigenomics provides insight into why certain people are more susceptible to obesity than others, even when following similar lifestyles. For example, variants in genes such as FTO (fat mass and obesity-associated gene), MC4R (melanocortin 4 receptor), and PPARG (peroxisome proliferator-activated receptor gamma) have been shown to influence energy intake, adipogenesis, and insulin sensitivity. As the field matures, nutrigenomics holds the potential to guide individualized dietary strategies that are more effective and sustainable than generalized dietary guidelines. Tailoring nutrition to an individual’s genetic makeup enables targeted prevention of obesity and related comorbidities, thereby improving long-term health outcomes and reducing the burden on healthcare systems. This article examines the molecular underpinnings of nutrigenomics in relation to obesity, highlighting key gene-nutrient interactions, and explores how precision nutrition can be leveraged to create actionable, personalized dietary interventions. Through clinical examples and case applications such as RxPREDICT gene-based reports, the article underscores the potential of nutrigenomics to transform the landscape of obesity prevention and personalized health management.
The primary characteristic of obesity is the accumulation of abnormal fat, which creates significant health risks. The World Health Organization defines overweight status as a BMI value of 25 or higher and obesity status as a BMI value of 30 or higher.1 The process of obesity develops when a person consumes more calories than their body uses for energy.2 The process of consuming more calories than your body needs leads to the storage of fat, resulting in weight gain over time. The impact of obesity on quality of life remained unrecorded until the 18th century despite its emergence as a worldwide public health problem. The first documentation of obesity-related morbid complications appeared during the early 20th century.3 The world currently has two billion adults and 340 million children who suffer from obesity, and this number is projected to double during the following decade. People who are overweight or obese experience discriminatory behaviors and rejection from their peers as well as prejudicial attitudes (e.g., “fat people are disgusting”) and negative stereotypical beliefs. The general public typically holds the view that obesity stems from poor lifestyle choices and unhealthy eating habits. The development of excess body weight is influenced by multiple factors, including metabolic rate, immune regulation, gut microbiota, adipose tissue browning, and genetic and epigenetic profiles.2 Genetic variation plays a crucial role in contributing to the development of obesity.4 Nutrigenomics is the scientific field that investigates how nutrients affect the genetic expression patterns of individual people.5 The examination of bioactive molecules' interactions with genes reveals their potential to modify gene expression through upregulation and downregulation, as well as activation and silencing of specific genes.4
An individual's genetic profile may influence susceptibility to obesity. Studies have identified genetic variations that participate in complex interactions between genes and dietary factors, influencing body composition and weight. In this regard, nutrigenetics, the study of how genetic diversity affects an individual's nutritional requirements, may advance knowledge of weight management and aid in the personalized dietary treatment of obesity.
A significant public health concern in both industrialized and developing nations, obesity is a complex and polygenic illness. Among the significant health conditions that contribute to morbidity linked to a higher incidence of obesity are cardiovascular disease, type 2 diabetes, non-alcoholic fatty liver disease, metabolic syndrome, and cancer.6 Possible evidence of the relationship between obesity and inflammatory diseases, which are linked to the genesis of cardiovascular disease, type 2 diabetes, and cancer, has emerged since the 1990s. The predominance of environmental factors (the so-called obesogenic environment) over genetic factors in explaining why obesity has reached epidemic proportions.7
Obesity can be classified as either polygenic, where the total number of SNPs determines an individual's weight through the combined influence of several genes (each of which has a negligible effect), or monogenic, where genetic variants in a single gene or a specific disease-related chromosomal region are the cause. Many genes linked to obesity regulate energy intake, lipid metabolism, adipogenesis, thermogenesis, adipocytokine production, and the activity of transcription factors. Polygenic obesity has a broad, complex, and non-deterministic genetic base. Clinical practice is complicated because many variations are dispersed across the genome and only contribute slightly to the onset of obesity. If adequately presented and comprehended, a collection of genetic variation information is required to describe prevention and therapy.8
The Human Genome Project marked a transformative period in health research, laying the foundation for personalized medicine and significantly impacting fields such as nutrition science. Nutritional genomics, also known as nutrigenomics, emerged from this, focusing on how individual genetic variations influence responses to nutrients and dietary components. This field aims to develop personalized nutrition plans that optimize health and prevent diseases based on an individual’s genetic profile. However, to make genome sequencing clinically useful, researchers must manage vast datasets and identify which genetic markers are most relevant to health. This involves advanced bioinformatics, data storage solutions, and tools for interpreting complex genetic data, particularly in relation to individual health outcomes. Nutrigenomics is now investigating how specific nutrients interact with genes and how these interactions can lead to personalized nutrition therapies, helping to prevent or manage conditions such as obesity, diabetes, and cardiovascular diseases.
International genome projects utilizing whole-genome sequencing have significantly expanded our understanding of genetic diversity, identifying numerous genetic variations, including single-nucleotide polymorphisms (SNPs), copy number variations (CNVs), and other structural variants, across diverse populations. These insights are foundational for nutrigenetics, which explores how genetic variants influence individuals' responses to specific nutrients and dietary patterns.9
Nutrigenetic research has identified genetic variants that interact with dietary factors to influence disease susceptibility, underscoring the intricate relationship between genes and nutrition. This knowledge significantly contributes to the prevention and management of chronic diseases. Single-nucleotide polymorphisms (SNPs), or more broadly, single-nucleotide variants (SNVs) that include common and low-frequency alleles, are among the most extensively studied genetic variations in precision nutrition. Many SNPs have been linked to chronic diseases through interactions with nutrient intake or dietary patterns (Table 1). For instance, polymorphisms in genes associated with taste perception, such as TAS1R2 (a sweet taste receptor) and CD36 (cluster of differentiation 36), have been linked to dyslipidemia in Mexican individuals consuming elevated levels of carbohydrates and fats, respectively. Common variants in genes involved in homocysteine metabolism, such as MTHFR (methylenetetrahydrofolate reductase) and MTR (methionine synthase), are associated with an increased risk of breast cancer in individuals with low intakes of folate, vitamin B6, and vitamin B12. Additionally, vitamin D status is influenced by various polymorphisms in vitamin D pathway genes, which can modulate its biological effects. Notably, SNPs in the VDR (vitamin D receptor) gene, which affect vitamin D availability, have been linked to an increased risk of osteoporosis in postmenopausal women with low calcium intake.10 Vitamin D plays a critical role in maintaining cardiovascular health. Historically, vitamin D deficiency has primarily been linked to skeletal issues, such as osteoporosis, osteomalacia, osteopenia, and increased risk of fractures. However, emerging evidence now suggests that insufficient vitamin D levels are associated with a broader array of non-skeletal disorders, including certain cancers, neurological disorders, metabolic diseases, and cardiovascular disease (CVD). The role of genetic factors in influencing individual responses to vitamin D supplementation is an area of active research, especially among specific populations such as postmenopausal women with overweight or obesity and individuals with obesity and type 2 diabetes. Genetic variations, or single-nucleotide polymorphisms (SNPs), in genes related to vitamin D metabolism and receptor activity can significantly affect how individuals respond to vitamin D supplementation. Studies have highlighted specific SNPs that may influence serum 25-hydroxyvitamin D [25(OH)D] levels after supplementation. For instance, a study involving 2,207 postmenopausal women with overweight and obesity identified the SNP rs11185644 in the retinoid X receptor alpha (RXRA) gene as being significantly associated with variations in the dose-response to vitamin D supplementation. This suggests that individuals with specific RXRA gene variants may experience different changes in serum 25(OH)D levels following vitamin D intake. In people with type 2 diabetes, particular SNPs in the vitamin D receptor (VDR) gene have also been linked to differing responses to vitamin D supplementation. These studies highlight the role of genetic variation in determining individual responses to vitamin D supplementation, particularly in populations with specific health conditions, such as obesity and type 2 diabetes. The presence of certain VDR and RXRA gene variants may modulate the effectiveness of vitamin D supplementation in influencing vitamin D levels and inflammatory or metabolic health markers. Analyzing complex genetic data to evaluate gene–diet interactions involve employing a variety of statistical approaches that address the high dimensionality and complexity of the data.15 Understanding these genetic differences could eventually lead to more personalized approaches to vitamin D supplementation, optimizing treatment based on an individual's genetic makeup.
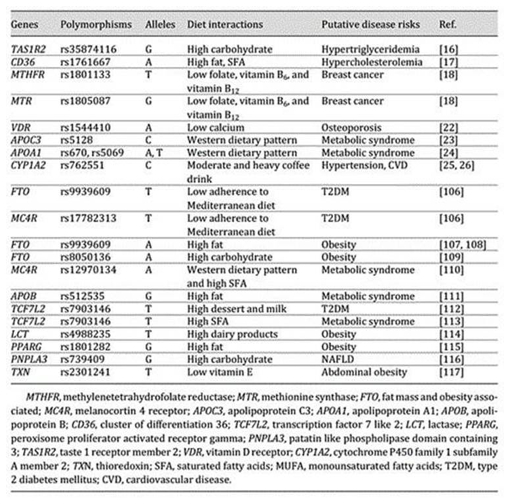
Table 1 SNPs linked to chronic diseases through interactions with nutrient intake or dietary patterns
|
Report summary |
Report summary |
Report summary |
|
Weight loss |
Weight loss |
Weight loss |
|
Report |
Rating |
Genes Associated |
|
Weight Loss Tendency |
Below Average |
FTO, TCF7L2, MTNR1B, PPARG, BDNF, ABCB11, PPARG |
|
Metabolism |
Above Average |
LEPR, CHRNA3, CRY2 |
|
Weight Regain |
Above Average |
FTO, PPARG, BDNF, NEGR1, TMEM18, KTCD15, GNPDA2 |
|
Satiety |
Well Below Average |
FTO |
|
FOOD |
FOOD |
FOOD |
|
Protein Utilization |
Normal |
FTO, LCT |
|
Fat Utilization |
Low |
PPARG, TCF7L2, APOA5, CRY2, MTNR1B, PPM1K |
|
Carb Utilization |
Enhanced |
IRS1, FGF21 |
|
Food sensitivity |
Food sensitivity |
Food sensitivity |
|
Gluten Sensitivity |
Low |
HLA-DQ |
|
Lactose Intolerance |
Likely |
MCM6 |
|
Caffeine Metabolism |
Slow |
Intergenic, AHR, RP11-10017.3-001, ARID3B, CYP1A1 |
|
Alcohol Sensitivity |
Moderately Sensitive |
ADH1C, ADG1B, ALHD2 |
|
Sweets Preference |
Above Average |
FGF21, SLCA2 |
|
Bitterness Sensitivity |
Likely |
TAS2R38 |
|
Impulse Control & Taste Preference |
Normal |
FTO |
|
NUTRIENTS |
NUTRIENTS |
NUTRIENTS |
|
Vitamin B9 (Folate) Tendency |
Normal |
MTHFR |
|
Vitamin B12 Tendency |
Normal |
FUT2 |
|
Vitamin B6 Tendency |
Low |
NBPF3 |
|
Vitamin A Tendency |
Normal |
BCMO1 |
|
Vitamin C Tendency |
Normal |
SLC23A1 |
|
Vitamin D Tendency |
Below Average |
GC, NADSYN1, CYP2R1 |
|
Vitamin E Tendency |
Above Average |
ZPR1, SCARB1, CYP4F2 |
|
Calcium Tendency |
Normal |
CASR, DGKD, GCKR, LINC00709, CARS, LOC105370176, CYP24A1 |
|
Copper Tendency |
Normal |
SMIM1, SELENBP1 |
|
Iron Tendency |
Normal |
TRF2, HFE, HFE, TMPRSS6 |
|
Magnesium Tendency |
Normal |
MUC1, ATP2B1, DCDC5, TRPM6, SHROOM3, MDS1 |
|
Phosphorus Tendency |
Normal |
ALPL, CSTA, IHPK3, PDE7B, C12orf4, IP6K3 |
|
Selenium Tendency |
Above Average |
DMGDH |
|
Zinc Tendency |
Normal |
CA1, PPCDC, LINC01420 |
|
Exercise |
Exercise |
Exercise |
|
Fat Loss Response to Cardio |
Normal |
ADRB2, LPL |
|
Fitness Response to Cardio |
Normal |
AMPD1, APOE |
|
Body Composition Response to Strength Training |
Enhanced |
NRXN3, GNPDA2, LRRN6C, PRKD1, GPRC5B, SLC39A8, FTO, FLJ35779, MAP2K5, QPCTL-GIPR, NEGR1, LRP1B, MTCH2, MTIF3, RPL27A, SEC16B, FAIM2, FANCL, ETV5, TFAP2B |
|
HDL Response to Cardio |
Below Average |
APOE |
|
Glucose Response to Cardio |
Normal |
PPARG |
|
Insulin Sensitivity Response to Cardio |
Below Average |
LIPC |
Table 2 Anonymous RxPREDICT report summary
|
Report summary |
Report summary |
Report summary |
|
Weight loss |
Weight loss |
Weight loss |
|
Report |
Rating |
Genes associated |
|
Weight Loss Tendency |
Normal |
FTO, TCF7L2, MTNR1B, PPARG, BDNF, ABCB11, PPARG |
|
Metabolism |
Above Average |
LEPR, CHRNA3, CRY2 |
|
Weight Regain |
Normal |
FTO, PPARG, BDNF, NEGR1, TMEM18, KTCD15, GNPDA2 |
|
Satiety |
Below Average |
FTO |
|
FOOD |
FOOD |
FOOD |
|
Protein Utilization |
Normal |
FTO, LCT |
|
Fat Utilization |
Normal |
PPARG, TCF7L2, APOA5, CRY2, MTNR1B, PPM1K |
|
Carb Utilization |
Enhanced |
IRS1, FGF21 |
|
Food sensitivity |
Food sensitivity |
Food sensitivity |
|
Gluten Sensitivity |
High |
HLA-DQ |
|
Lactose Intolerance |
Likely |
MCM6 |
|
Caffeine Metabolism |
Slow |
Intergenic, AHR, RP11-10017.3-001,ARID3B, CYP1A1 |
|
Alcohol Sensitivity |
Not Sensitive |
ADH1C, ADG1B, ALHD2 |
|
Sweets Preference |
Above Average |
FGF21, SLCA2 |
|
Bitterness Sensitivity |
Likely |
TAS2R38 |
|
Impulse Control & Taste Preference |
Normal |
FTO |
|
NUTRIENTS |
NUTRIENTS |
NUTRIENTS |
|
Vitamin B9 (Folate) Tendency |
Normal |
MTHFR |
|
Vitamin B12 Tendency |
Normal |
FUT2 |
|
Vitamin B6 Tendency |
Low |
NBPF3 |
|
Vitamin A Tendency |
Normal |
BCMO1 |
|
Vitamin C Tendency |
Normal |
SLC23A1 |
|
Vitamin D Tendency |
Below Average |
GC, NADSYN1, CYP2R1 |
|
Vitamin E Tendency |
Above Average |
ZPR1, SCARB1, CYP4F2 |
|
Calcium Tendency |
Normal |
CASR, DGKD, GCKR, LINC00709, CARS, LOC105370176, CYP24A1 |
|
Copper Tendency |
Normal |
SMIM1, SELENBP1 |
|
Iron Tendency |
Above Average |
TRF2, HFE, HFE, TMPRSS6 |
|
Magnesium Tendency |
Normal |
MUC1, ATP2B1, DCDC5, TRPM6, SHROOM3, MDS1 |
|
Phosphorus Tendency |
Above Average |
ALPL, CSTA, IHPK3, PDE7B, C12orf4, IP6K3 |
|
Selenium Tendency |
Normal |
DMGDH |
|
Zinc Tendency |
Normal |
CA1, PPCDC, LINC01420 |
|
Exercise |
Exercise |
Exercise |
|
Fat Loss Response to Cardio |
Normal |
ADRB2, LPL |
|
Fitness Response to Cardio |
Normal |
AMPD1, APOE |
|
Body Composition Response to Strength Training |
Enhanced |
NRXN3, GNPDA2, LRRN6C, PRKD1, GPRC5B, SLC39A8, FTO, FLJ35779, MAP2K5, QPCTL-GIPR, NEGR1, LRP1B, MTCH2, MTIF3, RPL27A, SEC16B, FAIM2, FANCL, ETV5, TFAP2B |
|
HDL Response to Cardio |
Enhanced |
APOE |
|
Glucose Response to Cardio |
Normal |
PPARG |
|
Insulin Sensitivity Response to Cardio |
Enhanced |
LIPC |
Table 3 Anonymous RxPredict report summary
In addition to macronutrient distribution, nutrigenomics can inform micronutrient intake. Certain genetic variations may increase the body's demand for specific vitamins and minerals that influence metabolic pathways involved in regulating weight. For instance, vitamin D status is linked to fat metabolism, and certain individuals may have genetic polymorphisms that reduce their response to dietary vitamin D, necessitating higher intake or supplementation. Similarly, individuals with variations affecting antioxidant pathways may benefit from higher intakes of nutrients, such as vitamins C and E, to reduce oxidative stress, which is linked to obesity and metabolic syndrome. Nutrigenomic insights can thus help refine dietary recommendations to address specific micronutrient needs that support weight control and overall metabolic health.11
Variants in lipid-related genes, such as APOC3 (apolipoprotein C3) and APOA1 (apolipoprotein A1) have also been shown to increase metabolic syndrome risk in individuals following a Western dietary pattern. Additionally, a variant in the CYP1A2 gene (cytochrome P450 family one subfamily A member 2) has been linked to a higher risk of hypertension and cardiovascular disease among moderate and heavy coffee drinkers. Research using genetic risk scores (GRS) has further examined the cumulative impact of SNPs on diet-related disease susceptibility. For example, macronutrient intake has been shown to influence the association between obesity GRS and increased adiposity. Interactions between saturated fat intake and an obesity GRS have been found to modify BMI in two American populations. Likewise, obesity GRS has been shown to interact with sugar-sweetened beverage and fried food consumption concerning BMI and obesity across several cohort studies.10
Gene-specific dietary plans for obesity
RxPREDICT is a company that offers genetic testing to create diet and exercise plans tailored to an individual’s genetic profile for personalized weight loss. Personalized nutrition (PN) represents a transformative approach in dietary science, where individual genetic profiles guide tailored dietary recommendations, thereby optimizing health outcomes and managing chronic diseases more effectively,12 they provide a report summary covering aspects like weight loss tendencies, metabolic rate, weight regain, satiety, and utilization of proteins, fats, and carbohydrates, as well as sensitivities to specific foods and nutrients. Each section is rated as above average, normal, or below average. For example, an anonymous report may show that an individual has a normal metabolic rate, meaning they are more likely to burn an average number of calories per day outside of physical activity. Your resting metabolism may change favorably in response to a balanced diet and regular exercise.
The RxPREDICT reports in tables 2 and 3 comprehensively show how an individual’s genetic profile influences their weight loss journey, food metabolism, and exercise response. For example, one individual’s results indicate a normal tendency for weight loss but below-average satiety, suggesting a genetic predisposition to feeling less full after meals, which could lead to overeating. Their above-average metabolism may counteract this by increasing daily caloric burn. In terms of macronutrient processing, they exhibit enhanced carbohydrate utilization but low-fat utilization, meaning a diet higher in complex carbohydrates and lower in fats may be more effective for weight management. Sensitivities, such as lactose intolerance and slow caffeine metabolism, also inform dietary restrictions and caffeine intake. On the exercise side, the individual exhibits a normal fat loss and fitness response to cardio, but an enhanced body composition response to strength training, suggesting that strength-focused routines may yield better results. The report highlights genes that influence these results. These insights highlight the potential of personalized plans, informed by genetic data, to improve outcomes in obesity treatment and overall health.
Gene-specific dietary plans for obesity
RxPREDICT is a company that offers genetic testing to create diet and exercise plans tailored to an individual’s genetic profile for personalized weight loss. Personalized nutrition (PN) represents a transformative approach in dietary science, where individual genetic profiles guide tailored dietary recommendations, thereby optimizing health outcomes and managing chronic diseases more effectively.12 They provide a report summary covering aspects like weight loss tendencies, metabolic rate, weight regain, satiety, and utilization of proteins, fats, and carbohydrates, as well as sensitivities to specific foods and nutrients. Each section is rated as above average, normal, or below average. For example, an anonymous report may indicate that an individual has a normal metabolic rate, meaning they are more likely to burn an average number of calories per day, regardless of physical activity. Your resting metabolism may change favorably in response to a balanced diet and regular exercise.
In conclusion, nutrigenomics holds immense potential for preventing and managing obesity by offering precise, personalized dietary recommendations. By understanding the genetic factors that contribute to weight gain and metabolic health, healthcare providers can tailor dietary plans to each individual’s unique needs, thereby increasing the likelihood of successful weight management and long-term health benefits. As research and technology in nutrigenomics continue to advance, the integration of genetic insights into dietary planning may become a cornerstone of preventive health strategies, reducing the global burden of obesity and empowering individuals to achieve optimal health through targeted, science-based nutrition.13-15
None.
There is no conflict of interest.

©2025 Gupta, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.