MOJ
eISSN: 2471-139X


Review Article Volume 3 Issue 3
1Department of Cell Biology and Anatomy, Rosalind Franklin University of Medicine and Science, USA
2Division of Biomedical Statistics and Informatics, Mayo Clinic, USA
Correspondence: Michael P Sarras, Department of Cell Biology and Anatomy, Rosalind Franklin University of Medicine and Science, 3333 Green Bay Road, North Chicago, IL 60064, USA
Received: February 19, 2017 | Published: March 20, 2017
Citation: Sarras MP, Leontovich AA. Hydra’s ECM/integrin/matrix metalloproteinase network: comparative analysis of hydra and vertebrate ECM signaling systems based on current genomic and EST databases. MOJ Anat Physiol. 2017;3(3):96–105. DOI: 10.15406/mojap.2017.03.00094
The fundamental and critical functions of the ECM to metazoans explain why we see its appearance in ancient Taxa such as Cnidaria that represents one of the earliest groups with a defined tissue organization and an ECM. A model system for understanding the mechanisms involved in Cell/ECM interactions during developmental processes is the Cnidarian, Hydra (diverged some 500million years ago) whose use in developmental biology dates back to the studies of Trembley in 1774. As discussed in this review, ECM does not stand alone, but rather is part of a complicated system that we refer to as the Extracellular matrix/Integrin/Matrix Metalloproteinase Network (ECM/Integrin/MMP Network). This network not only provides structural support for tissues, but also functions as a signal transduction platform that regulates cell physiological and developmental processes. The purpose of the review is to show the highly conserved nature of the ECM/Integrin/MMP Network throughout the metazoans and does so by comparing what we know of the vertebrate network to that of the Hydra network.
The review is organized to discuss
Keywords: extracellular matrix, hydra, integrins, MMPS, development, regeneration
ECM, extracellular matrix; PG, proteoglycan; GAG, glycosaminoglycan; SLRP, small leucine rich proteoglycan; FACIT, fibril associated collagens with interrupted triple helices; MACIT, membrane associated collagens with interrupted triple helices; MULTIPLEXIN, multiple triple helix domains and interruptions/endostatin producing collagens; MMP, matrix metalloproteinase
Approximately 500million years ago early metazoans diverged with defined tissue layers associated with an extracellular matrix (ECM).1 As seen in modern vertebrates, the ECM not only provides a structural foundation for the tissues, but also exists as a signaling platform to orchestrate a wide spectrum of cues to surrounding cells that guides cell differentiation, morphogenesis and general cell physiological processes.2,3 It does this by integrating its structural components and associated molecules with cell surface receptors (e.g. Integrins) that bind to motifs within the various matrix components to transmit signals to the ECM-associated cells.2‒4 These processes are facilitated by enzymes [Matrix Metalloproteinases (MMP)] that not only remodel the ECM during morphogenesis but also function to release signaling molecules from the matrix for subsequent binding to cell surface receptors. This entire integrated system can be referred to as the Extracellular Matrix/Integrin/Matrix Metalloproteinase Network (ECM/Integrin/MMP Network).
The critical and fundamental function of metazoan ECM explains why we see its appearance in ancient Taxa such as Cnidaria that represents one of the earliest eukaryotic systems with a defined tissue organization.1 Cnidarians diverged some 500million years ago and are composed of an outer ectoderm and inner endoderm with an intervening acellular material called the mesoglea that is now known to be a true ECM. A Cnidarian model system for understanding the mechanisms involved in Cell/ECM interactions during developmental processes is Hydra. Hydra is a member of the Hydrozoan Class of Cnidaria whose use in developmental biology dates back to the studies of Trembley in 1774.5 Based on biochemical, cellular, molecular studies focused on understanding the components and function of various macromolecules of the mesoglea of Hydra, we now have a basic understanding of the role of ECM in an early divergent metazoan that predated and forecasted what we see in the later more complicated vertebrate systems.1,6
This review focuses on our understanding of the ECM/Integrin/MMP Network as reflected in Hydra. Its purpose is to show the highly conserved nature of the ECM/Integrin/MMP Network throughout the metazoans and does this by comparing what we know of the vertebrate network to that of the Hydra network based on current NCBI
Genomic databases7(NCBI Hydra Genome Database: https://www.ncbi.nlm.nih.gov/genome/?term=Hydra) and
EST databases8 (NCBI Hydra EST Databases: https://www.ncbi.nlm.nih.gov/nucest. The current review expands on a 2014 review by Tucker & Adams9 that focused on adhesion molecules of Cnidarians.
The review will focus on
The structure of hydra and its ECM
As indicated, Cnidarians are the earliest group of metazoans with a defined tissue organization (ectoderm and endoderm) and an ECM. A tree of Taxa divergence is shown in Figure 1. This tree of Taxa with a “Common Tool Kit” for the ECM/Integrin/MMP Network indicates that Cnidaria containing Hydra arose some 500million years ago and represents one of the earliest metazoan life forms with an ECM.1,6
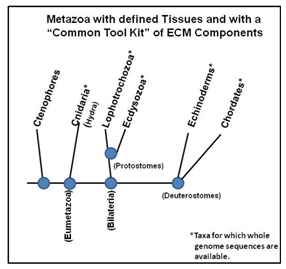
Figure 1 A hieratical tree of metazoans with a “Common Tool Kit” for the ECM/Integrin/MMP Network. Those Taxa with an asterisk have complete genome databases available.
The basic structure of Hydra’s body wall is shown in Figure 2. As indicated, the entire body wall of Hydra (from the foot process, through the body column, and apically to the tip of head tentacles) is organized as an outer ectoderm and inner endoderm with an intervening ECM, termed the mesoglea by early invertebrate biologists (Figure 2, top image). Based on molecular and cellular studies, Hydra’s ECM is itself organized as two basal laminas (adjacent to the two epithelial layers) with an intervening interstitial-like matrix composed of fibrillar collagens [such as Hcol-I]10 as shown in Figure 2, bottom image.
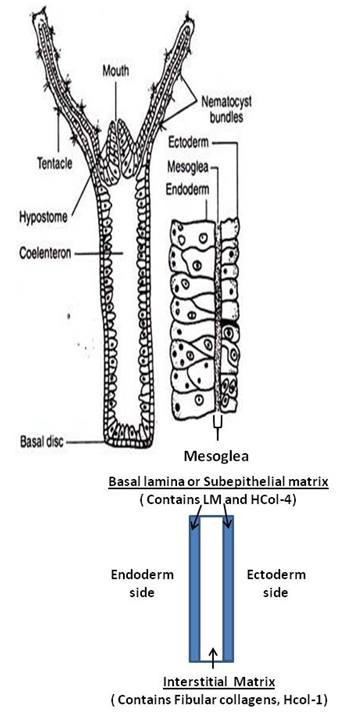
Figure 2 Hydra body plan formed from an epithelial bilayer with an intervening extracellular matrix (ECM). Hydra exists as a gastric tube with a mouth and several tentacles at the head pole and a peduncle and a basal disk at the foot pole (top diagram). The entire body wall of Hydra (from the tip of the tentacles to the basal disk) is organized as an epithelial bilayer with an intervening ECM along the entire longitudinal axis of the organism. Hydra’s ECM is structured as two sub epithelial zones (i.e. basal lamina matrix) with an intervening central fibrous zone (i.e. interstitial matrix). A simplified drawing of the mesoglea (extracellular matrix) is shown in the bottom image. Hydra laminin and Type IV Collagen are localized to the two sub epithelial zones of the matrix (basal lamina) while Hydra fibrillar collagens (e.g. Hcol-I) are localized to the central fibrous zone or interstitial matrix.
Initial approaches to the analysis of hydra ECM (prior to 1980)
Invertebrate biologists termed the extracellular material (an acellular matrix) between the two epithelial cell layers of Hydra’s body wall as the “mesoglea”. Later electron microscopy techniques identified a broad spectrum of structural components within this mesoglea. It was described as an amorphous matrix of low density containing fine fibrils ranging from 5 to 50nm in diameter.11‒13 Both ectodermal and endodermal cells were seen to send cellular processes from their basal surfaces into the mesoglea to form cell-cell contacts in the matrix.11,12,14
Studies by Davis & Haynes15 were performed on both contracted and relaxed Hydra to analyze the structure of mesoglea under these two conditions. They established that the inter-epithelial matrix was 0.5 to 2.0mm in diameter with it being most thick in the body column and most thin in the tentacles.15 Ultrastructural analysis15 found that mesoglea had three ultrastructural components:
Fibrils were of three types:
Additional studies by Shostak16 found that isolated Hydra mesoglea was highly elastic and had the ability to be stretched to twice its normal length. The mesoglea would immediately retract to its original length when tension was released.
In the 1970’s the first biochemical studies were performed on isolated mesoglea. Studies by Barzansky & Lenhoff17 & Barzansky et al.,18 utilized a combination of gel chromatography, amino acid analysis, and thin layer chromatography for sugar moiety analysis in conjunction with radiotracer precursor-labeling to probe the structural components of Hydra mesoglea. They found that Hydra mesoglea had biochemical characteristics similar to vertebrate basal lamina. They also concluded that the amino acid profiles suggested the presence of collagens. Experiments using lathyritic agents known to cause reduced cross-linking of collagens, indicated a role for these macromolecules in Hydra regenerative processes (e.g. head and foot regeneration).17
In summary, these early studies established that Hydra mesoglea had many of the properties of vertebrate extracellular ECM in terms of both structural and biochemical characteristics. This set the foundation for later studies beginning in the 1980’s that would utilize a combination of structural, biochemical, cellular, and molecular approaches to establish the true nature of the Hydra ECM.
Initiation of the hydra ECM project incorporating structural, biochemical, cellular, and molecular approaches to analysis of hydra ECM (after 1980)
As indicated, a comprehensive analysis of Hydra mesoglea began in the 1980’s with the initiation of the Hydra ECM Project.10,19‒21,21‒26 The strategy of this project was multifaceted and its objective was to understand the structure, composition, and function of the Hydra mesoglea using modern techniques.
It was designed to
The results of the Hydra ECM Project established that Hydra mesoglea was reflective of a vertebrate ECM with many of the matrix components and associated molecules seen in Chordates. The basic structure and general molecular composition of Hydra ECM is depicted in Figure 2. It was found that Hydra ECM had the components and characteristics of both basal lamina and interstitial matrix. In addition, ECM associated molecules such as matrix cell surface receptors (e.g. integrin-like)26 and matrix remodeling enzymes such as matrix metalloproteinases (MMPs) were also identified.25 From the one original Hydra MMP25 that was cloned and functionally characterized, we can now identify sixteen hydra MMPs based on NCBI databases as will be discussed later in the review. All of these ECM-associated components were critical to Hydra cell physiological processes (e.g. cell differentiation, cell migration, and shape changes) as well as regeneration and general morphogenesis. The reader is referred to a detailed review of these findings published in 2012.26 Rather than repeat what has been previously reviewed, we will now extend those studies by comparing our current understanding of vertebrate ECM and associated molecules (see recent reviews)2,3 to that of Hydra using current NCBI genomic and EST databases. We refer to the ECM and its associated molecules as the ECM/Integrin/MMP Network because we now know that ECM is not simply a structural entity, but also a complex signaling entity that controls a broad range of cellular and developmental functions.2,3,27‒29
Composition of the vertebrate ECM/Integrin/MMP network
ECM components: The matrix macromolecules of the vertebrate ECM generally fall into three categories. These three categories include:
Protein Class and Members |
Protein Class and Members (Cont.) |
Proteoglycans |
Glycoproteins. |
Extracellular Proteoglycans: |
Fibronectin (FN): |
Hyalectans |
FN, only 1 form |
Aggrecan |
|
Versican |
|
Neurocan |
Fibrillin: |
Brevian |
One form |
Small Leucine-rich Proteoglycans: |
Fibulin/Hemicentin: |
Decorin |
One form |
Biglycan |
|
Fibromodulin |
SPARC: |
Lumincan |
One form |
Pericellular-basement membrane Proteoglycans: |
Thrombospondin: |
Perlecan |
One form |
Agrin |
|
Cell Surface (CS) Proteoglycans: (Can also serve as CS binding Proteins) |
Laminins (LN) |
Syndecans |
16 Trimeric chains made up of three of the following chains to form a single heterotrimeric laminin: Five a chains (a 1-5)Three b chains (b 1-3) Three g chains (g 1-3) (A laminin made up of a1, b1, & g1 chains referred to as Laminin-1,1, 1) |
Glypicans |
|
CSP G4 |
|
Betaglycans |
|
Phosphacan |
|
Collagens. |
|
Fibrillar Collagens: |
|
I, II, III, V, XI, XXIV, and XXVII |
|
Network-forming Collagens: |
|
IV, VIII, and X |
|
FACITs: |
|
IX, XII, XIV, XVI, XIX, XX, XXI, XXII |
|
MACITs: |
|
XIII, XXVII, XXIII, XXV |
|
Anchoring Fibrils: |
|
VII |
|
Beaded-filament-forming Collagens: |
|
VI, XXVI, and XXVIII |
|
Multiplexins |
|
XV and XVIII |
Table 1 Vertebrate extracellular matrix (ECM) protein components of the EMC/integrin network.
Note: *Italicized ECM components with Hydra Analogs (Table 4).
The proteoglycan group is typically composed of a core protein with covalently attached glycosaminoglycan chains (GAGs).30,31 GAGs are highly negatively charged heteropolysacchrides that are composed of repeating disaccharides units. There are six types of GAGs that are typically synthesized in the Golgi apparatus for subsequent conjugation to the core proteins. The GAG, Hyaluronan (HA) is a-typical in that it is synthesized on the cell surface and seen without a conjugated core protein.32,33
For the purposes of this review, we will focus on the core proteins that are classified into four groups to include the:
The largest group is the extracellular group made up of
Hyalectans2 (HA and lectin binding PGs) contain: Aggrecan (cartilage-associated), Versican (expressed in various tissues and important in cell adhesion, migration, and inflammation), and Neurocan & Brevican (both expressed in the vertebrate brain). The SLRP group2 includes: decorin, biglycan, fibromodulin, and lumican. Decorin typically attaches to collagens through non-covalent interactions and forms a linker molecular for growth factors. Biglycan is expressed in many tissues and in vertebrates has an important role in postnatal bone growth via interactions with growth factors. Fibromodulin is widely distributed in connective tissues and has a role in collagen fibrillogenesis. Lumican is typically seen in mesenchymal tissues and its binding to collagen maintains proper fibril spacing. It also binds to growth factors and regulates development by doing so. The Pericellular proteoglycans are seen in association with various tissues such as the epithelium.2 They function in a variety of processes such as angiogenesis. They are composed of the perlecans and agrins. Perlecan is seen in both vertebrates and invertebrates and functions in cell regulation via its ability to bind to various ligands. Agrin is similar to perlecan and functions in cell regulation processes such as clustering of acetylcholine receptors. Cell surface proteoglycans are either transmembrane proteins or GPI-anchored proteins.2 Two common molecules of this group are syndecan and glypican; both of which are involved with cell regulation via growth factor interactions. Others in this group include: CSPG4, betaglycan, and phosphacan; all three of which regulate cell behavior through interaction with cell surface receptors and/or growth factors. The final group of PGs is the Intracellular Proteoglycans which are not associated with the ECM, but rather are found within the secretory compartment of cells (e.g. serglycin).2
The most common fibrous protein of the body is collagen which is found in both interstitial and basement membrane ECM.34 Collagens are made up of twenty eight different types, comprise up to 30% of the proteins of the vertebrate body, and are seen throughout the animal kingdom. The collagens are classified into seven different categories, based on common domain homology and function. These categories include: fibrillar collagens, network-forming collagens, fibril-associated collagens with interrupted triple helices (FACITs), membrane-associated collagens with interrupted triple helices (MACITs), anchoring fibrils, beaded-filament-forming collagens, and multiple-triple-helix domains and interruptions)/endostatin-producing collagens (MULTIPLEXINs) as listed in Table 1. Fibrillar collagens are common in a variety of tissues and provide tensile strength to these tissues. They are typically triple helical molecules that are extensively post-translationally modified in order to form proper fibrillar structure and this involves association with other ECM components such as decorin and biglycan.34 They also function in cell processes such as cell migration, adhesion, blood vessel formation, tissue repair, and tissue development. Network-forming collagens are typically found in basement membranes (basal lamina) to include the renal filtration barrier. These collagens assemble in combination with a number of ECM components such as the laminins and nidogens following discharge from the cell. The FACITs interact with fibrillar collagens and function to link fibers together in the matrix.34 MACITs are transmembrane proteins with a C-terminal domain that is composed of collagen regions and non-collagen domains (NC-domains).34 They function as cell surface receptors affecting a variety of cell processes such as cell adhesion and migration. Anchoring fibrils link basement membrane components to the interstitial matrix components that support the basement membrane. Beaded-filament–forming collagens function as additional linkers between ECM components34 and MULTIPLEINs also serve as linkers in the ECM and when cleaved by matrix associated enzymes, can trigger cell shape changes induced by Integrin receptor signal transduction pathways affecting cytoskeletal networks within cells.34
A broad array of non-collagen glycoproteins is also critical to ECM structure and function. These include: fibronectin,35 laminins,36,37 SPARC,2 fibulins2 and nidogens2 to name only a few. As with the collagens, these molecules can form interlacing networks within the ECM and upon which many ECM associated molecules can bind for structural integrality and signaling functions. All of these ECM components are listed in Table 1. Those vertebrate ECM components with Hydra analogs are highlighted in yellow.
Integrin components: Integrins are heterodimeric receptors made of Alpha (a) and Beta (b) subunits.38 They serve as signal transducers between the ECM and adjacent cells. They function along with other ECM binding proteins such as Discoidin-domain-receptors and CD44. They also function in cell differentiation, cell shape changes, and in ECM assembly, to name but a few of their functions. A list of the Integrins and other ECM-associated cell surface binding proteins are listed in Table 2. Those receptors with Hydra analogs are highlighted in yellow.
Protein Class and Members |
Integrins. |
Dimeric Receptors: |
Each integrin receptor is made up of both an α and β chain. |
There are 18 α subunits (α1-18) and 8 β subunits (β1-8) to form the various α/β Integrins. |
Discoidin Domin Receptors |
DDR1 |
DDR2 |
CD44 |
A single trans membrane glycoprotein that functions as a cell surface receptor for HA |
Table 2 Vertebrate extracellular matrix receptors.
Note: *Italicized receptors indicate those with Hydra analogs (Table 5).
Matrix metalloproteinase components: The most significant classes of enzymes that specifically degrade vertebrate ECM are the metalloproteinases. These enzymes were initially isolated in studies designed to determine the biochemical nature of the molecules responsible for tadpole tail resorption during metamorphosis to the adult frog.39 These metalloproteinase fall into two families to include:
The matrix metalloproteinases (MMPs)4 and the disintegrin and metalloproteinase with thrombospondin motifs family, called the ADAMTSs/ADAMs.4 Because genomic and EST studies have identified many MMP genes and mRNAs for Hydra, but have identified fewer ADAMTSs/ADAMs in Hydra whose function(s) has not been evaluated, this review has focused only on the MMP family of metalloproteinases. For the reader’s reference, Hydra ADAMTSs/ADAMs from Genomic and ETS data bases include: ADAMDEC1-like, gene ID#: 105846829; ADAMTS6-like, accession#: XP 002168102; ADAMTS20-like, accession#: XP 002166940; ADAMTS-like, accession#: XP 002169841; and ADAM-like, accession#: XP 002161708.
Since the original identification of tadpole tail matrix metalloproteinase [later determined to be MMP3 based on cDNA studies]40 a total of twenty four vertebrate MMPs have been identified through a combination of biochemical and molecular approaches.4 A summary of these vertebrate MMPs is shown in Table 3. As will be discussed in Section 5, sixteen of these vertebrates MMPs have Hydra analogs and these MMPs are italicized in Table 3. The table not only lists the classification of these vertebrate MMPs based on structural characteristics, but also lists their ECM substrates based on biochemical analyses. These MMPs also have other substrates such as growth factors and other MMPs (as utilized during the activation of MMPs).4 It should be noted that human MMPs have critical functions in a broad spectrum of processes involving morphogenesis, cell physiology, and abnormalities related to disease.4
MMP by Structural Group |
ECM Substrates |
MMPs with Basic MMP Structure |
|
MMP 1 |
Collagens I, II, III, VII, & X; gelatins; aggrecan; entactin; tencasin; perlecan. |
MMP 3 |
Aggrecans; decorin; gelatins; fibronectin; laminin; collagens III, IV, IX & X; tenascin; perlecan. |
MMP 8 |
Collagens I, II & III; gelatins, aggrecan |
MMP 10 |
Aggrecan; Fibronectin; laminin; collagens III, IV, and V |
MMP 11 |
Fibronectin; laminin; aggrecan; gelatins. |
MMP 12 |
Elastin; aggrecan; Fibronectin; Osteonectin; laminin; nidogen. |
MMP 13 |
Collagens I, II, III, IV, IX, X, & XIV; aggrecan; dibronectin; tenascin; SPARC/oseteonectin; laminin; perlecan. |
MMP 18 |
Collagens (Xenopus). |
MMP 19 |
Nidogen-1 and general ECM components such as in Matrigel. |
MMP 20 |
Tooth enamel. |
MMP 21 |
Not known, but involved in breast cancer and left/right symmentry during embryogenesis. |
MMP 27 |
Fibronectin; laminin; gelatins and/or collagens. |
MMP 28 |
Casein. |
MMPs with Fibronectin-Domain Inserts |
|
MMP 2 |
Gelatins; Collagens IV, V, VII, X, & XI; fibronectin, laminin; elastin; aggrecan. |
MMP 9 |
Gelatins; Collagens III, IV, & V; aggrecan; elastin; entactin; vitronectin. |
Membrane-Bound MMPs |
|
Trans membrane: |
|
MMP 14 |
Collagens I, II, III; gelatina; aggrecan; fibronectin; laminin; fibrin. |
MMP 15 |
Fibronectin; laminin; tenascin; nidigen; aggrecan; perlecan. |
MMP 16 |
Collagens III; fibronectin; gelatin. |
MMP 24 |
Dermatin; Chondroitin sulfate; fibronectin; Fibrin; gelatin. |
GPI-anchored: |
|
MMP 17 |
Gelatin; fibrinogen. |
MMP 23 |
Not known. |
MMP 25 |
Gelatin; Collagens IV; fibrin; fibronectin; laminin. |
Minimal Domain MMPs |
|
MMP 7 |
Aggrecan; gelatin; fibronectin; laminin; elastin; entacin; collagen IV; tenascin; decorin. |
MMP 26 |
Gelatin; Collagens IV; fibronectin; fibrinogen; vitronectin. |
Table 3 Vertebrate MMPs with ECM Substrates.
Note: *Italicized MMPs have potential Hydra analogs (Table 6)
Protein Class and Members |
Protein Class and Members (Cont.) |
[With NCBI Gene Identifier # (*GI #:); Gene ID# (*GID#): or Accession # (*AC#:)] |
|
Proteoglycans. |
Glveoproteins. |
Extracellular Proteoglycans: |
Fibronectin (FN): |
Versican *GI#: 50349993 |
Novel Hydra Protein with a FN Type!!! domain containing RGD motifs *GI#: 74125001 |
Pericellular-basement membrane Proteoglvcans: |
|
Perlecan *AC#: XP_002168817 |
Fibrillin: |
Agrin *GI#: 31083785 |
One form *AC#: XP_001630852 |
Cell Surface (CS) Proteoglvcans: |
Fibulin/Hemicentin: |
One form *AC#: XP_002154934 |
|
Syndecan *GIN: 105844043 |
|
Collagens. |
Thrombospondin: |
Fibrillar Collagens (w/chain type): |
One form *AC#: XP_002164610 |
I α *GI#: 68409940; α2 *Gl#: 56539382; α3 *GI#: 68399093 |
|
ll α1 *GI#: 57832129 |
|
III α1 *GI#: 68410130 |
|
V α1*GID#: 105844122; α1 *GI#: 47547182; α2 *GI#: 46963722; α1 *GI#: 68409389 |
|
XI α1 *GI#: 68409434; α2 *GI52859140 |
|
XXIV α1 *GI#: 68385074 |
|
Network-forming Collagens (w/chain type): |
Laminins (LN) [(α), (β) and (ϒ) Chains]: |
IV α1 *Gl#: 684109;: α2 *Gl#: 74120560; |
Alpha Chains: |
α4 *Gl#: 68409671; α5 *Gl#: 68409584; |
LN α1 *GID: 100197510; *AC#: DN815114.2 |
α6 *Gl#: 68401718 |
LN α2 *GIN: 101235032 |
X α1 *Gl#: 68396063 |
LN α3 *GID#: 101241783 |
FACITs (w/chain type): |
Beta Chains: |
IX α1 *Gl#: 68380272; α2 *GI# 68381420 |
LN β1 *GID#: 100202556 |
XII α1 *Gl#: 47548498 |
LN β3 *GID#: 101236303 |
XIV α1 *GID#: 105849478; α1 *Gl# 654460808 XVI |
Gamma Chains: |
XVI α1 *Gl#: 68378121 |
LN ϒ1 *GID#: 100198511; *AC#: DT613962.1 |
MIACITs (w/chain type): |
|
XXVII α1 *GI#: 68391894 |
|
Beaded-filament-forming Collagens (w/chain type): |
SPARC: |
VI; α1 *Gl#: 74125949; α6 *GID#: 105848864 |
One form *AC#: XP_002167014 |
Multiplexins (w/chain types): |
|
XV *Gl#: 47143009 |
|
XVIII α1 *GI#: 68381740 |
|
Table 4 Hydra Extracellular Matrix (ECM) Proteins of the ECM component of the ECM/Integrin Network
Composition of the hydra ECM/Integrin/MMP Network based on current NCBI hydra databases
As previously indicated, the data that follows regarding Hydra’s ECM/Integrin/MMP Network was obtained from current NCBI databases derived from both genomic and EST studies. Genomic sequence databases are based on the studies of Chapman et al.7,41
(NCBI Hydra Genome Database https://www.ncbi.nlm.nih.gov/genome/?term=Hydra) and EST databases are based on the studies of Wenger & Galliot8 (NCBI Hydra EST Database: https://www.ncbi.nlm.nih.gov/nucest).
ECM components: The original Hydra ECM Project established the basic structure of Hydra ECM and also identified ECM-associated molecules such as Hydra cell surface ECM binding proteins (e.g. Integrin-like receptors) and MMPs.25 The single molecule cloning approach used in those studies was laborious but in combination with antibody, cellular, and biochemical studies it produced a good overview of Hydra ECM and its associated components. However, with the advent of genomic and EST studies, we now have a much broader database to analyze. Such an analysis yielded the Hydra ECM components listed in Table 4. This takes the number of Hydra ECM structural components from four (laminin a1 and b1 chains, collagen IV, and a single fibrillar collagen) based on the original studies, to thirty one identified components based on the results from current genomic and EST databases.
One conundrum that may be explained from this analysis is the role fibronectin (with its many RGD motifs) plays in Hydra. Studies by Stidwill & Christen,42 as well as other investigators,10,19,22,23,43 have previously established that RGD peptides and antibodies to fibronectin were effective in blocking cell migration and morphogenesis in Hydra; however, various cDNA screening methods failed to yield a Hydra fibronectin cDNA clone. From the current NCBI database studies it is clear that a novel Hydra protein with fibronectin type III domains (domains known to contain RGD peptide motifs) exists in Hydra which offers an explanation to the fibronectin-like results that were previously published. The presence of RDG motifs in collagen IV’s NC domain (Non-collagen domain) may also have a role in the fibronectin-like processes previously reported in Hydra.
Integrin components: Original Hydra ECM studies provided evidence for ECM binding proteins,19,22,23 but did not identify Hydra integrin cDNA clones. Analysis of current genomic and EST databases however, clearly show a broad range of both Alpha (a) and Beta (b) submit chains as shown in Table 5. This expands the biochemical and functional studies previous published and indicates that Hydra has at least eight integrin subunits to form various heterodimeric receptors for induction of ECM-cell signal transduction pathways in the organism.
Chain Subunit Class and Members |
|
Integrins. |
Gene ID#. |
Alpha chains (α): |
|
Alpha-4 chain (α4): |
101239034 |
Alpha-5 chain (α5, αV): |
100208443 |
Alpha-7 chain (α7): |
101237856 |
Alpha-8 chain (α8): |
100202151 |
Beta chains (β): |
Beta-2 chains (β2) |
100123119 |
Beta-3 chains (β3) |
101238114 |
Beta-PS-like chain (β-PS) (Hydra Drosophila analog) |
100205067 |
Beta-pat-like chain (β-pat-3) (Hydra C. elegans analog) |
105844257 |
Table 5 Hydra Extracellular Matrix (ECM) Receptors of the ECM/Integrin Network.
In total, these studies indicated that Integrins emerged in parallel with ECM formation during the divergence of early metazoan Taxa such as the Cnidarians.
Matrix metalloproteinase components: As part of the Hydra ECM Project,19‒21,26,44 additional studies were initiated to determine if Hydra expressed MMPs as seen in vertebrates. Using mammalian derived MMP probes, screening of Hydra cDNA libraries yielded the isolation of a Hydra MMP, named HMMP.25 Sequence analysis of the full length HMMP cDNA found that the molecule had sequence similarity with a number of vertebrate MMPs.25
Protein expression of the HMMP clone achieved a functional enzyme that was then analyzed to determine its biochemical characteristics.25 The activated enzyme (following cleavage of the Pro-region) was found to be 46 KDa while the deduced full-length enzyme (with its Pro-peptide) was 55.4 KDa. HMMP had an optimal pH of 6.5-7.5 and this activity was inhibited by GM6001, a known inhibitor of mammalian MMPs. Like mammalian MMPs, HMMP was inhibited by matlistatin and recombinant human TIMP-1.25 In keeping with known MMP hydrolytic properties, HMMP could degrade isolated Hydra ECM and this degradation was blocked by GM6001.25 Analysis of the hydrolysis products of Hydra ECM incubated with HMMP found that some of these components reacted with antibody to the b1-laminin chain; indicating that HMMP likely had laminin as one its substrates. In vitro analysis determined that HMMP was not able to hydrolyze mammalian type I collagen, type IV collagen, laminin, or fibronectin; while these same ECM components could be degraded by human MMP3. This indicates a high degree of species hydrolytic specification for HMMP.
Gene expression and functional studies indicated an up-regulation of HMMP mRNA during regenerative processes such as head and foot regeneration. Blockage of HMMP activity with GM6001 during induced regeneration, inhibited regenerative processes. Convergent studies using antisense thio-oligo nucleotides were also found to block regenerative processes; thereby supporting the GM6001 studies.25 Since the cloning and functional characterization of Hydra HMMP, only a few articles have been published that focus on the detailed characterization of Hydra MMPs. This includes the work of Wenger et al.45 in 2014 dealing with injury-induced immune responses in Hydra,45 Buzgariu et al.46 in 2015 dealing with multi-functionality and plasticity of Hydra epithelial cells,46 and a recent 2016 article dealing with the effect of UV on foot duplication in regenerating Hydra and the role of HMMP in that process (Table 6).47
Genomic Analyses |
EST Analyses |
||
MMPs with the Basic MMP structure |
MMPs with the Basic MMP structure |
||
Gene ID#: |
Accession#: |
||
MMP3 |
101239608 |
MMP1 |
CN767751.1 |
MMP11 |
100202175 |
MMP3 |
CV863180.2 |
MMP18 |
101234333 |
MMP8 |
CX833843.2 |
MMP19 |
101241586 |
MMP12 |
DY451023.1 |
MMP27 |
100203699 |
MMP13 |
DY452652.1 |
MMPs with Fibronectin-Domain Inserts |
MMPs with Fibronectin-Domain Inserts |
||
Gene ID#: |
Accession#: |
||
MMP2 |
100199972 |
MMP2 |
DT614350.1 |
Membrane-Bound MMPs |
Membrane-Bound MMPs |
||
Trans-membrane: |
Gene ID#: |
Trans-membrane: |
Accession#: |
MMP 14 |
100200849 |
MMP 24 |
DN813429.2 |
MMP 16 |
105847473 |
||
MMP 24 |
100201778 |
||
GPI-anchored: |
Gene ID#: |
||
MMP 17 |
105845208 |
||
MMP 25 |
100200988 |
||
Minimal Domain MMPs |
Gene ID#: |
Minimal Domain MMPs |
Accession#: |
MMP 7 |
100197160 |
MMP 7 |
DN138495.2 |
Table 6 Hydra MMPs identified from Genomics and EST Analyses
In total, these studies indicated that MMPs emerged in parallel with ECM and Integrin formation during the divergence of early metazoan Taxa such as the Cnidarians.
From available studies and databases, it is clear that the ECM/Integrin/MMP Network emerged early during the formation of metazoans as reflected from this analysis of Hydra’s ECM and ECM-associated components. This likely reflects the critical function of the ECM/Integrin/MMP Network in metazoans. This network’s functions encompass complex processes involving not only the structural integrity of tissues, but the dynamic nature of ECM to regulate basic cell processes such as; cell differentiation, morphogenesis, regeneration, and development in general. As a final note, the reader should be aware that ECM/Integrin/MMP Network studies have expanded to include not only the transmission of signals from the ECM to the cell, but also the concept of the “Integrin Adhesome Network” which includes all internal cellular pathways triggered by the externally originating ECM signals.48,49
The authors wish to thank those who have developed the genomic and EST databases that have allowed a more in depth study of the relationship of Hydra ECM to vertebrate ECM.
Author declares that there is no conflict of interest.

©2017 Sarras, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.