eISSN: 2379-6367


Review Article Volume 1 Issue 1
Division of Pharmaceutical Sciences, Long Island University, USA
Correspondence: Vikas Sehdev, Division of Pharmaceutical Sciences, Long Island University, Arnold & Marie Schwartz College of Pharmacy and Health Sciences, HS 608, Brooklyn, NY - 11201, USA , Tel 718 488 1447, Fax 718 780 4586
Received: November 03, 2014 | Published: November 18, 2014
Citation: Betzu JM, Jasani N, Desai B, et al. Selective targeting of M3 muscarinic receptors: an opportunity for improved treatment of upper gastrointestinal carcinomas. Pharm Pharmacol Int J. 2014;1(1):1-5. DOI: 10.15406/ppij.2014.01.00001
Upper Gastrointestinal (GI) Carcinomas are one of the most morbid forms of cancers that are inherently resistant to chemotherapeutic regimens. The G-protein coupled receptors (GPCRs) are the largest family of cell surface receptors that have recently emerged as significant modulators of cancer cell growth, survival, and metastasis. Owing to the normal physiologic role of muscarinic receptors, a major sub-class of GPCRs, many epithelial cells undergoes proliferation when exposed to acetylcholine. Since majority of the cancers are epithelial in origin, cancer cells frequently take over the philological machinery associated with muscarinic receptors and undergo uncontrolled proliferation, avoid cell death, and exhibit invasion and migration. The M3 sub-type of muscarinic receptors is over expressed in tumors of different types. In this review we focus on the oncogenic signaling mechanisms activated by the M3 receptors. The manuscript highlights the importance of M3 receptor as a potent therapeutic target for selective treatment of Upper GI Carcinomas. We also reported, for the very first time, that M3 receptor mRNA is significantly over expressed in Upper GI Carcinoma tissue samples. In addition, we also showed that M3-specific anti-muscarinic agents have considerable anticancer activity in Upper GI Cancer cells. This review serves as foundation for future studies delineating the anti-tumor effect of M3-specific anti-muscarinic agents as single agents or in combination in Upper GI Carcinoma.
Keywords: upper gastrointestinal cancer, m3 muscarinic receptor, solifenacin, darifenacin, oxybutynin, microarray
PI3K, phosphoinositide 3-kinase; Akt or PKB, protein kinase B; cAMP, cyclic adenosine monophosphate; ATP, adenosine triphosphate; NFκB, nuclear factor kappa b; APC, adenomatous polyposis Coli; GEO, gene expression omnibus; GI, gastrointestinal; EGFR, epidermal growth factor receptor; HER-2, human epidermal growth factor receptor 2; AURKA, aurora kinase A; IP3, inositol triphosphate; PLC, phospholipase C; DAG, diacylglycerol; MAPK, mitogen activated protein kinasE; RTKs, receptor tyrosine kinase; MMP9, matrix metalloproteinase 9; CHO, chinese hamster ovary; PGE2, prostaglandin E2; NFκB, nuclear factor kappa B; COX-2, cyclooxygenase-2; 5-FU, 5 fluorouracil; DMSO, dimethyl sulfoxide
Upper Gastrointestinal (GI) Cancers (i.e. cancers of both stomach and esophagus) are the third most frequently diagnosed form of cancer world-wide.1 Latest GLOBOCAN 2012 global epidemiological data indicate that Upper GI Carcinoma accounts for approximately 1.12million deaths annually making it the second most morbid form of cancer.1 In addition, as per the Surveillance, Epidemiology, and End Results (SEER) data published by National Cancer Institute the incidence and mortality rates for gastric cancers in US have reduced marginally, however, there has been no change in these parameters for esophageal cancers for the last three decades.
Upper GI Carcinomas an aggressive malignancy characterized by inherent resistance to current chemotherapeutic regimens, high rates of disease recurrence, tumor metastasis and poor patient survival.2‒6 Multiple potent oncogenic molecular pathways driven by the epidermal growth factor receptor (EGFR), human epidermal growth factor receptor 2 (HER-2), v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (K-RAS), and Aurora kinase A (AURKA) pathways are constitutively overactive in Upper GI Carcinomas.7‒11 Therefore, development and characterization of novel chemotherapeutic regimens based on the molecular makeup of Upper GI Carcinomas are urgently needed to reduce patient mortality and morbidity.
The G-protein coupled receptors (GPCRs) are a super-family of receptor proteins that account for the largest number of cell surface receptors expressed by any human cell.12 GPCRs modulate multiple signaling pathways that play a vital role in maintaining normal cellular physiology and homeostasis. GPCRs mediatepotentsignaling mechanisms that govern normal physiological responses associated with cell proliferation, survival and motility.13 Unfortunately, aberrant and deregulated activity of GPCRs has been observed in various types of cancers and is associated with malignant transformation due to overactive oncogenic signaling mechanisms resulting in tumor growth, drug resistance, angiogenesis and metastasis.13 The Muscarinic Cholinergic Receptors (mAChRs) are one of the five major families of GPCRs and consist of five distinct subtypes (M1, M2, M3, M4, and M5).12 The M1, M3 and M5 mAChR sare coupled to Gαq subunit of G-proteins and on activation they stimulate phospholipase C (PLC) enzyme that further breaks down phosphatidylinositol 4,5-bisphosphate (PI ) into diacylglycerol (DAG) and inositol triphosphate (IP3) resulting in increased intracellular levels of calcium.14,15 Elevated intracellular levels of calcium can activate multiple growth, survival and migration promoting signaling mechanisms. The activated Gαq subunit stimulates PLC-β that further activates the mitogen activated protein kinase (MAPK) pathway.16 In addition βγ subunit of the G-proteins has been indicated to activate receptor tyrosine kinase (RTKs), Src and PI3K that mediate growth and survival promoting downstream signaling mechanisms. The M2 and M4 mAChRs are inhibitory in function and interact with Gαi/Gα0 subunit of G-proteins. Their activation results in reduced synthesis of cAMP from cellular ATP within the cells.15 Studies conducted in various animal models have shown that M1, M3 and M5 mAChRs are expressed in normal epithelial cells of the Upper GI tract.17‒19 Majority of the cancers are derived from epithelial cells that have undergone malignant transformation. Therefore, most of them exhibit aberrant muscarinic signaling in the form of over expressed mAChR sand/or enhanced activation by acetylcholine (ACh) secreted from autocrine and paracrine sources.
The M3 mAChR is the subtype majorly over expressed in lung, skin, colon, gastric, pancreatic (subtype not identified), breast, ovarian, prostate and brain cancers.20‒30 The M3 mAChR has been shown to induce multiple signaling pathways associated with cellular growth, survival, inflammation, angiogenesis, invasion and migration in different types of cancer cells. Some of the key oncogenic cellular mechanisms activated by M3 receptors in cancer cells include the EGFR/MAPK pathway (growth and proliferation), PI3K/Akt pathway (pro-survival and anti-apoptosis), β-catenin/Wnt pathway (invasion and migration) and NFκB pathway (pro-inflammatory and chemotactic).
The proliferative MAPK pathway and the pro-survival PI3K/Akt pathway are frequently activated by cell surface localized RTKs. EGFR is one such RTK that istransactivatedby M3 receptors through paracrine acetylcholine signaling mechanisms in colon cancer cells.31 Transactivation of EGFR by M3 receptor stimulates the MAPK pathway resulting in colon cancer cell proliferation.32 In small-cell lung cancer (SCLC) inhibition of M3 receptor with Darifenacin (an M3-specific antagonist) reduced the activity of MAPK pathway and suppressed overall tumor growth in vitro and in vivo.33 Specific inhibition of M3receptor with 4-DAMP (an M3-specific antagonist) and M3 receptor knock down suppressed phosphorylation of MAPK in MCF-7 breast cancer cells.27 In addition, M3 receptor mediated activation of P3K/Akt pathway has also been indicated to play a vital role in inducing growth and proliferation of astrocytoma cells.34
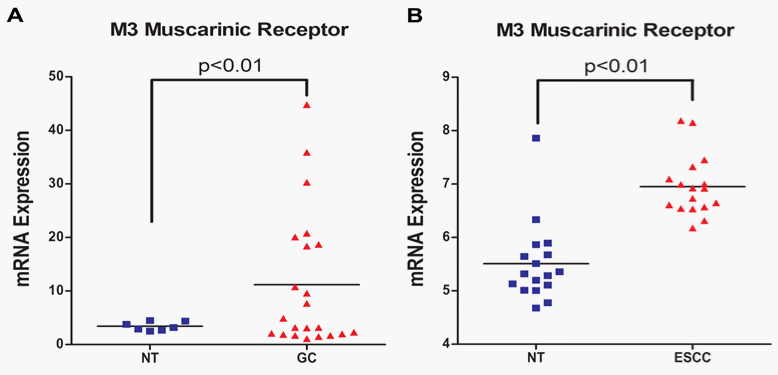
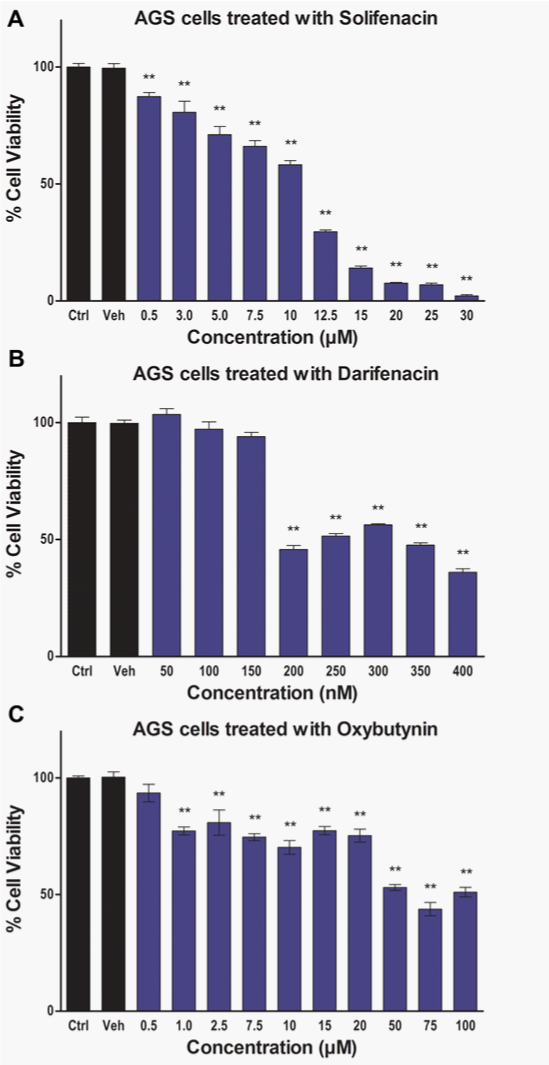
The M3 receptor signaling is particularly pronounced in the lungs where M3 receptor expression has been correlated with tumor metastasis and poor survival rates in patients with non–small-cell lung cancer (NSCLC).20 The M3 receptors are indicated to induce invasion and migration of NSCLC cancer cell lines by enhancing the expression and activity of matrix metalloproteinase 9 (MMP9) through the PI3K/Akt pathway.20 In melanomas, M3 receptor expression has also been correlated with invasion and migration.22 The Wnt/β-catenin signaling pathway plays an important role in inducing invasion and metastasis of cancer cells. The regulatory effect of M3 receptor on Wnt/β-catenin signaling pathway was determined by Raufmann et al. in Apcmin/+ mice that were knocked out for M3 receptor gene.35 Their data indicated that M3 receptor knockout Apcmin/+ mice exhibited reduced activity of β-catenin as evidenced by decreased nuclear localization. Furthermore, M3receptor can mediate drug resistance since it has been shown to prevent induction of P53 pro-apoptotic protein in Chinese hamster ovary (CHO) cells following treatment with a DNA-damaging agent.36 The M3 receptors have also been implicated in promoting angiogenesis by inducing nitric oxide synthesis, prostaglandin E2 (PGE2) production and VEGF expression in breast cancer cells.37‒39 Chronic Inflammation is an important precursor for carcinogenesis. The nuclear factor kappa B (NFκB) pathway plays a critical role in regulating expression of genes mediating inflammation. Carbachol induced M3 receptor activation has been associated with increased activation of NFκB pathway in human astrocytoma cells and up-regulation of cyclooxygenase-2 (COX-2) enzyme mediated PGE2 synthesis in colon cancer cells.40,41 Together, NFκB and PGE2 can mediate chronic inflammatory response resulting in sustained cellular damage that can ultimately induce malignant transformation. These observations suggest that M3 receptor function is frequently up-regulated in cancer cells and it can play an important role in the overall progression of various types of cancers. However, there is paucity in scientific literature with respect to the status of M3 receptor expression and function in human Upper GI Carcinomas. At present the status of muscarinic receptor expression in normal human esophageal epithelial cells and cancer cell lines is unknown. In addition, there is only one study published by Kodaira et al.25 that demonstrated the presence of M3 receptors in various human gastric cancer cell lines.25 Therefore, to further establish the clinical relevance of M3 receptor expression for the treatment of Upper GI Carcinomas we first investigated the presence of mRNA for various muscarinic receptors (M1-M5) in tumor tissue samples from patients. We mined the publically available NCBI GEO database for microarray data associated with normal and cancerous Upper GI tissue samples. The microarray data (Figure 1A) (Figure 1B) indicate that mRNA expression of M3 receptor is significantly high in both gastric (normal gastric tissue: 3.429±0.3068, N=7; gastric cancer: 11.17±2.689, N=22; p<0.01)42 and esophageal (normal esophageal tissue:5.510±0.1795, N=17; esophageal cancer: 6.953±0.1373, N=17; p<0.01)43 cancers. Our data analysis did not show any significant increase in the mRNA expression for M1, M2, M4 and M5 receptors in the aforementioned microarray data sets. We also used various M3 receptor specific anti-muscarinic agents (solifenacin, darifenacin, and oxybutynin) and determined their effect on Upper GI Cancer cell viability. Our novel data exhibits significant anticancer activity by solifenacin, darifenacin and oxybutynin in AGS Upper GI Cancer cells (Figures 2A‒2C).
The aberrant GPCR signaling network plays a pivotal role in activating multiple oncogenic mechanisms involved in cancer development and progression. The M3 mAChR is one of the major GPCRs that are up-regulated indifferent types of cancers and it induces growth, survival, drug resistance, angiogenesis, invasion, migration and inflammation associated signaling mechanisms. The microarray data indicate that M3 receptors are up-regulated in Upper GI Carcinomas and the cell viability data shows that M3-specific anti-muscarinic agents inhibit Upper GI Cancer cell viability. At present, promising clinically approved M3-specific anti-muscarinic drugs are available that are well tolerated and exhibit minimal side effects. Since Upper GI Carcinomas exhibit resistance to multiple chemotherapeutic agents it is possible that a combination regimen consisting of a first line chemotherapeutic agent like Cisplatin, Docetaxel, or 5 Fluorouracil (5-FU) with M3-specific anti-muscarinic agents will show enhanced antitumor activity. Overall, based on M3 receptor function, expression and availability of M3-specific anti-muscarinic agents we suggest that future studies should be conducted to determine the therapeutic value of selective targeting of M3 receptors for the treatment of Upper GI Carcinomas.
Cell culture and pharmacologic reagents
AGS Upper GI Cancer cell line was maintained as a monolayer culture in F12 Media (Gibco) cell culture medium supplemented with 10% (v/v) FBS (Gibco). Solifenacin and Oxybutynin stock solutions were prepared in water and further diluted in cell culture media for cell viability assay. Darifenacin stock solution was prepared in dimethyl sulfoxide (DMSO).
MTT cell viability assay
AGS cells were seeded at 2000 cells/well into 96-well plates. The plated cells were treated with Solifenacin (0.5–30μM), Darifenacin (50–400 nM), or Oxybutynin (0.5–100μM) and incubated for 72 hours at 37°C and 5% CO2 level in an incubator (Panasonic Healthcare Co.). Subsequently, 25μL of MTT reagent (5mg/mL in phosphate buffered saline pH 7.2 or PBS) was added into each well and the plates were incubated for 4 hours. The violet colored formazan crystals formed in each well were dissolved in 100μL of DMSO. The absorbance of dissolved formazan crystals was measured at 540nm in a plate-reader (BioTek Instruments, Inc.). The intensity of the signal is a measure of overall cellular metabolic activity and an indicator of viable cells present at the end of the treatment.
Statistical analysis
Data are presented as means±SEM. All cell viability experiments were done in triplicates. One-way ANOVA with Tukey post hoc analysis was used to show statistical difference between control and treatment groups. Statistical analyses were done with Graph Pad Prism 5 software (Graph Pad Software, Inc.). For microarray RNA expression analysis T-test was done to determine statistical difference between normal tissue and tumor tissue samples. The p value of ≤0.05 was considered statistically significant. Statistically significant differences are marked in the Figures; ** p<0.01.
None.
Author declares that there is no conflict of interest.

©2014 Betzu, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.