eISSN: 2379-6367


Research Article Volume 8 Issue 5
University of Arkansas at Little Rock, Center for Molecular Design and Development, Department of Chemistry, Arkansas, USA
Correspondence: Jerry A Darsey, Center for Molecular Design and Development, Department of Chemistry, 2801 South University Avenue, Little Rock, Arkansas, 72211, USA
Received: September 30, 2020 | Published: October 26, 2020
Citation: Masarweh N, Darsey JA. Computational modelling of drugs for Alzheimer’s disease (AD) and applications on artificial neural network systems (ANN); NETS. Pharm Pharmacol Int J. 2020;8(5):312-316. DOI: 10.15406/ppij.2020.08.00310
Alzheimer’s disease (AD) is an irreversible and progressive disease that affects neurons and their connections in parts of the brain, specifically the hippocampus and entorhinal cortex. The purpose of this research is to modify current medications of Alzheimer’s disease with the use of computational modelling. The modifications are concluded to improve the half maximal inhibitory concentration (IC50) value which is the concentration needed for the drug to inhibit a specific biological function. Drug design throughout this research has been done on the computational modelling software Gaussian 09. The expected modified IC50 values are predicted using two methods. First, the functional graph methods utilizing the energies and the experimentally measured IC50 values producing correlations that result in predicted IC50 values for the modified drug molecules. The second method involves using an artificial neural network system NETS to predict the IC50values of modified drug molecules. Four modified drug molecules resulted in promising outcomes in which the IC50 values were improved with a value of one order of magnitude and higher. The data obtained shows that computational modelling can be a novel time-saving and significant step for drug discovery.
Keywords: Alzheimer’s disease, drug design, IC50, computational modelling, ab initio, artificial intelligence, artificial neural networks
AD, Alzheimer’s disease; ADHD, attention deficit hyperactivity disorder; ADD, attention deficit disorder; NMDA, (N-methyl-D-aspartate); AI, artificial intelligence; ANN, artificial neural network; IC50, half inhibitory concentration; pIC50, negative log of IC50; DFT, density functional theory; HOMO, highest occupied molecular orbital; LUMO, lowest unoccupied molecular orbital; QSAR, quantitative structure-activity relationship models; PES, potential energy surfaces; A.U., atomic units
Alzheimer's disease (AD) is the most common cause of dementia, it accounts for 60-80% of dementia cases.1 It is a progressive neurodegenerative disease that mainly targets memory and other important mental functions. In the United States, there are approximately 5.8 million patients that are currently diagnosed with Alzheimer’s, and the numbers are projected to triple to 16 million by 2050.2
Molecular and cellular understanding of the mechanism of this disease helps achieve a better approach to an effective therapy. The two main proteins responsible for Alzheimer’s are the excess production and accumulation of Tau and Beta-Amyloid proteins, but Tau and Beta-Amyloid proteins are not the only factors causing AD, other factors may include; the vascular system which may not be supplying enough blood and nutrients to the brain, the brain may lack the glucose to power its’ activities, or due to chronic inflammation that causes microglial cells (they are a specialized population of macrophages that are found in the central nervous system, they remove damaged neurons and infections and are important for maintaining the health of the central nervous system) to fail to do their job in clearing away debris and astrocytes which in return reacts to distressed microglial.3
The half-maximal inhibitory concentration (IC50) is a quantitative measure of a drug’s potency. It can indicate the amount of drug needed to inhibit a certain biological process by half, thus it can be used as a measure of potency of an antagonist drug in research. The relationship between the increase in the IC50 value and the potency of the drug is inversely proportional, the lower the IC50 value meaning the lower amount/concentration of a certain drug is needed to inhibit a specific biological function thus the better the potency. Generally, IC50 values are expressed in molar concentrations, either µM or nM but are sometimes converted into pIC50 scale to ease calculations (pIC50 = -log(IC50)).
The current approved medications to treat the symptoms of Alzheimer’s disease include cholinesterase inhibitors Donepezil, Rivastigmine, Galantamine and the NMDA (N-methyl-D-aspartate) receptor antagonist Memantine. Another medication, Methylphenidate, which is commonly used for narcolepsy and ADHD (attention deficit hyperactivity disorder) patients, has also been approved to treat apathy in AD patients.
These drugs provide symptomatic relief but are not effective as a cure. Drug discovery has been directed, in the last 10 years, to developing ‘disease modifying drugs’ hopefully able to counteract the progression of AD. Because in a chronic, slow progressing pathological process, such as AD, an early start of treatment enhances the chance of success. It is crucial to have biomarkers for early detection of AD-related brain dysfunction, preferably before clinical onset.4
Computational modelling is a theoretical field of chemistry that uses molecular orbitals to optimize, evaluate and help better understand the chemical processes and reactions that take place. Computational modelling can calculate energy differences between various states of the molecules to help understand mechanisms of reactions at the molecular level and can calculate the systematic stability of chemical systems. It has become a useful way to investigate materials that are too complex or difficult to find or very expensive to purchase. It also helps chemists make predictions before running the actual experiments so that they can be better prepared for predicting properties. The basis for much of computational modelling is the Schrödinger equation (a linear partial differential equation that shows the wave function or state function of a quantum-mechanical system), this is because this equation models the molecules and atoms with mathematics. It is a very useful tool for determining the momentum energy, transition structures, electronic structure determinations, frequency calculations, protein calculations, potential energy surfaces (PES), rate constants for chemical reactions (kinetics), thermodynamic calculations- heat of reactions and energy of activation.5
There are several types of computational modelling methods used by chemists, including, semi-empirical methods, molecular mechanics, molecular dynamics, quantum mechanics/molecular mechanics and lastly, ab initio methods which the Gaussian 09 computer modelling software is classified under. Ab initio methods are based on quantum chemistry. Ab initio means “from first principles”, meaning that only physical constants are inputted into an ab initio calculation. These ab initio methods work by solving the Schrödinger equation to yield helpful information such as energies and electron densities.
Gaussian 09 is a computer program usually used by chemical engineers, chemists, biochemists, and physicists. It utilizes fundamental laws of quantum mechanics to predict such parameters as energies, molecular orbitals, molecular structures, spectroscopic data (NMR, IR, UV, etc.) and many more. One technique employed is the density functional theory (DFT) method for the determination of molecular electronic structure. The uniqueness of DFT method is that the energy is expressed in terms of total one-electron density instead of a wave function.
The advantage of using a program such as Gaussian 09 is the ease of modifying molecules as needed, the vast quantum mechanical functions are easily performed, the wide variety of output given such as total energy, highest occupied molecular orbitals (HOMOs) and lowest unoccupied molecular orbitals (LUMOs), in addition to many others such as dipole moment, the angles and special distance between the molecules. All of which is of great value for the calculations intended for this work. Briefly, the structure of the molecule is inputted into the program where each atom must have a bond length, bond angle and dihedral angle, which locates each atom spatially in three dimensions. The minimum energy geometry of the molecule is then determined using the Gaussian 09 program.
The next step is to use the Artificial Intelligence (AI) software, NETS, in which the inputs into the AI program are the LUMOs, HOMOs, total energy and dipole moment.7 The specific AI software used is an Artificial Neural Network (ANN), which is a computing technique that mimics the biological neural network of animal brains. The program works by learning to correlate the input values to the output value/values. In this work, the output value that is being correlated is the pIC50 value of each molecule.6,7
With the constant need to develop novel drugs for treating diseases, the urge to constantly improve current on market medications for diseases such as Alzheimer’s disease is an area of continuous research. Typically, the process of preclinical trials, where possible new medications are tested in the laboratory on animals can take time from several months up to a year or more to conduct. The method used in this research helps to narrow the candidates for preclinical and clinical trials. Our group can model hundreds to thousands of possibilities in the time that would take to do one trial.
Five drug molecules that are currently FDA approved to treat symptoms of Alzheimer’s disease were modified. The modifications were done using different functional groups which depended on several factors including crystallography of the target proteins responsible for AD (beta amyloid and tau proteins), the results of previous research on similar drug molecules and some of the modifications were based on trial and error bases. Some of these modifications included adding hydrogen, fluorine, and methyl, trifluoromethyl and ammonia groups. A detailed list of the modifications is discussed in Table 1.
|
Modification No. |
Atom/Molecule Added |
|
1 |
Substitution of terminal methyl (-CH3) with Fluorine atom (F) |
|
2 |
Substitution of terminal Hydrogen (-H) with Fluorine atom (F) |
|
3 |
Substitution of terminal Hydrogen (-H) with Chlorine atom (Cl) |
|
4 |
Substitution of terminal methyl (-CH3) with Chlorine atom (Cl) |
|
5 |
Substitution of terminal methyl (-CH3) with carboxylic acid group (COOH) |
|
6 |
Substitution of terminal methyl (-CH3) with Trifluoromethyl group (CF3) |
|
7 |
Substitution of terminal Hydrogen (-H) with Ammonia atom (NH3) |
|
8 |
Substitution of Oxygen (O) with Sulfur atom (S) |
|
9 |
Substitution of terminal Hydrogen (-H) with Hydroxy group (OH) |
|
10 |
Substitution of terminal Hydrogen (-H) with Methoxy group (OCH3) |
|
11 |
Addition of Hydroxy (-OH) to Benzene ring |
|
12 |
Substitution of terminal Hydrogen (-H) with ethanol (CH2OH) |
|
13 |
Substitution of terminal Hydrogen (-H) with propanol ( CH2CH2CH2OH) |
|
14 |
Substitution of terminal Hydrogen (-H) with butanol (CH2CH2CH2CH2OH) |
|
15 |
Substitution of terminal Hydrogen (-H) with propionic acid (CH2CH2CHCOOH) |
Table 1 List of modifications on the original five drug molecules
The first method to calculate the predicted value of pIC50 was using functional graphs. Functional graphs are a method used to produce linear correlations between two variables. The data used to plot the functional graphs consist of the experimental pIC50 values vs. the total energy (calculated from Gaussian 09). Different pairs of data were used to produce different functional graphs, such as using the experimental pIC50 value vs. energy, using the inverse of the pIC50 value vs. energy, using the inverse of both the pIC50vs. energy, using the logarithm of pIC50 value vs. energy, using the logarithm of the pIC50 vs. the inverse of energy and using the pIC50 value vs. the inverse of energies. These plots were produced for all 15 sets of modifications. The linear functional graphs produce plots, from which we can calculate the predicted pIC50 values.
The second method which is a more rigorous technique uses artificial neural network (ANN) programming utilizing artificial intelligence (AI). The software NETS has been used for this purpose. This software uses back propagation algorithm. In machine learning, it is a widely used algorithm in training feed forward neural networks. Back propagation techniques are useful because they can efficiently compute the gradient of the loss function with respect to the weights of the network for a single input-output example. Unlike other techniques where direct computation of the gradient with respect to each weight individually.
The back propagation algorithm works by computing the gradient of the loss function with respect to each weight using the chain rule, computing the gradient one layer at a time, iterating backward from the last layer to avoid redundant calculations of intermediate terms in the chain rule.8 For more details see reference (8). The first step towards using AI to predict the pIC50 values is to prepare the input data to the ANN. NETS software utilizes a sigmoidal function to make its predictions. The data entered must be normalized between (0) and (1). The reason for normalization of data is due to the use of a sigmoidal function with limits between (0) and (1). For practical reasons we normalized between (0.05) and (0.95), using equation (1).
(1)
where:
xi: normalized value
x: variable (the numbers from data sets available).
Li: lowest number in data set series.
Hi: highest number in data set series.
Cross-validation techniques are used to validate the model by training on (n-1) data points and predicting the nth data point. This process is repeated until all (n) data points have been validated. This technique is performed by training on four values and predicting the fifth using ANN. A cross-validation linear plot consisting of experimental pIC50 values vs. the predicted pIC50 values is plotted. An r2 value as close to 1 as possible is desired. Training the neural network on a set of given data is an integral step to allow the machine learning to predict the pIC50 values for the modified drug molecules. This computational procedure has no limit as to how many modified molecules can be modelled.
Results
After the molecules have been modified and optimized on Gaussian 09, their energies, dipole moment, and HOMO-LUMO gap has been identified, it is now possible to construct functional correlation graphs to help us correlate the data obtained with the original data from literature in order to predict pIC50 values for the modified drug molecules. For the predicted pIC50 values, the higher the value predicted the more chance there is for the modified drug molecule to act as a better inhibitor of proteins responsible for causing AD. In this case, modifications 14 & 15 have been chosen as promising candidates for further research since they have resulted in enhanced values of IC50 (higher potency) for the corresponding drug molecules.
From the sample data in Table 2, multiple functional graphs have been plotted to find the best correlations between these data sets, which resulted in the best fit of inverse total energy vs. pIC50 values. By inserting information for Donepezil from Table 2 into the linear equation (2) extracted from Figure 2, we obtain the value of (x) which is the predicted pIC50 for the modified drug Donepezil.
|
Modified Drug Molecules |
pIC50 in Vivo (experimental) |
Total Energy (calculated) [A.U.] |
Inverse pIC50 in Vivo (experimental) |
Inverse Total Energy [A.U.] |
Log pIC50 in Vivo (experimental) |
|
Donepezil |
8.77 |
-1338.59 |
0.11402 |
-0.00075 |
0.943 |
|
Galantamine |
6.27 |
-1168.28 |
0.15949 |
-0.00086 |
0.797 |
|
Memantine |
5.26 |
-757.202 |
0.19011 |
-0.00132 |
0.72 |
|
Rivastigmine |
7.43 |
-919.68 |
0.13459 |
-0.00109 |
0.87 |
|
Methylphenidate |
7.08 |
-824.285 |
0.14124 |
-0.00121 |
0.85 |
Table 2 Example of data collected for plotting a functional correlation graph for modification no.14*
*Refer to Table 1 for modification no.14.
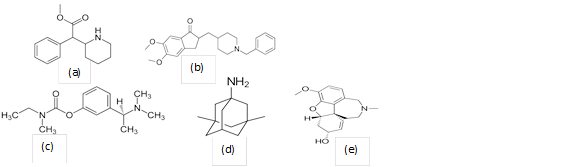
Figure 1 Chemical structure of: (a) Methylphenidate, (b) Donepezil, (c) Galantamine, (d) Rivastigmine, (e) Galantamine.
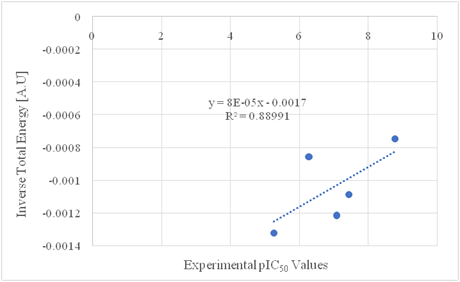
Figure 2 Functional correlation between inverse total energy [A.U.] and experimental pIC50 value for modification 14.
(2)
where,
Using the same procedure for modification 14, we obtained the predicted pIC50 value for modification 15, using Figure 3. Figure 4 shows the results of cross validation between the predicted pIC50 values from the NETS program vs. the experimental data. Table 3 shows a comparison of data between the values from the functional graphs method and values predicted using the AI method. Figure 5 shows a correlation plot of predicted pIC50 values obtained from AI method vs. calculated values predicted from functional graph method.
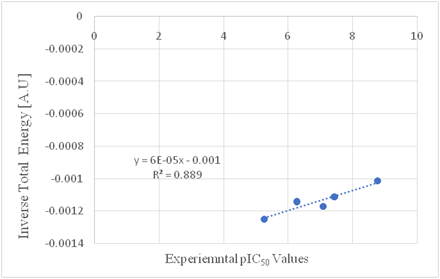
Figure 3 Functional correlation between inverse total energy [A.U.] and experimental pIC50 value for modification 15.
|
Modification |
pIC50 from functional graph |
pIC50 value from AI |
Percent Error [%] |
|
Donepezil |
11.91 |
11.87 |
0.33 |
|
Galantamine |
10.55 |
10.47 |
0.76 |
|
Memantine |
4.74 |
4.82 |
-1.69 |
|
Rivastigmine |
7.66 |
7.54 |
1.57 |
|
Ritalin |
6.09 |
6.15 |
-1.04 |
Table 3 Comparison of predictions made using functional graph method and the AI method
The purpose of this study was to find an innovative method to modify drugs used to treat symptoms of Alzheimer’s Disease by achieving a lower IC50 (higher potency) value of these drug modifications. The data obtained above shows promising results for the use of computational modelling for drug design. Four drug molecules; Donepezil, modifications 14 & 15 and Galantamine, modifications 14 &15 showed the greatest increase in potency (highest pIC50 values). The experimental pIC50 values for Donepezil and Galantamine are 8.77 and 6.27, respectively. Their pIC50 value after modification 14 (refer to Table 1) are 11.91 and 10.55, respectively. For modification 15 the values are 9.79 and 7.63, respectively. This demonstrates that medium chained alcohols and carboxylic acids produce the most significant impact on the enhancement of pIC50 values. Furthermore, the maximum percent error between the values predicted using functional methods and the ones from AI, (Table 3) was 1.57 percent. This illustrates that computational modelling coupled with artificial intelligence can be an effective tool for performing drug design.
The results obtained show success in the prediction of the modified pIC50 values. Computational modelling using ab initio molecular orbital methods and AI is a powerful approach to performing drug design. This approach can be used to model hundreds or thousands of drug modifications in the time it would take to perform only a few modifications using traditional experimental methods.
None.
None.
Authors declare that there is no conflict of interest.

©2020 Masarweh, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.