eISSN: 2379-6367


Research Article Volume 9 Issue 4
1Pharmaceutical Chemistry department, College of Pharmacy, Umm Al-Qura University, Saudi Arabia
2Clinical research center, The Executive Administration of Research and Innovation, King Abdullah Medical City (KAMC) in Holy Capital, Saudi Arabia
Correspondence: Sharaf E Sharaf, Pharmaceutical Chemistry Department, College of Pharmacy, Umm Al-Qura University, Makkah, Saudi Arabia, Tel +966532660411
Received: July 22, 2021 | Published: August 9, 2021
Citation: Sharaf SE. Biochemical computational therapeutic approach towards the discovery of natural product non-covalent inhibitors of SARS-CoV-2 RNAdependent RNA polymerase. Pharm Pharmacol Int J. 2021;9(4):160-169. DOI: 10.15406/ppij.2021.09.00341
COVID-19 remains to be a disastrous pandemic chaotically affecting our lives to this day. This disease is caused by the new coronavirus, SARS-CoV-2. Like other coronaviruses and RNA viruses, SARS-CoV-2 replication and transcription largely relies on the activity of RNA-dependent RNA polymerase (RdRp). This enzyme produces a complementary RNA molecule from a template one. The necessity of this enzyme for the viral lifecycle has made it an attractive target for combating many RNA viruses. The famous RdRp inhibitor remdesivir has been granted use for hospitalized COVID-19 patients. In this work, we investigate a library of natural products with the aim of discovering new RdRp inhibitors for potential therapeutic purposes. The study comprises extensive biochemical docking and molecular filtration of the database where the resultant hits were evaluated for other molecular properties and pharmacophoric features. Two hit compounds were discovered to be potential therapeutic RdRp inhibitors, Sennoside B and Ginsenoside B2. The metabolites of these compounds were also predicted to investigate whether they would possess extended activity against the enzyme.
Keywords: COVID-19, SARS-CoV-2, RdRp, Sennoside B, Ginsenoside B2
On March 11th, 2020, the World Health Organization (WHO) declared the SARS-CoV-2-induced disease, namely COVID-19, a pandemic of international concern https://www.who.int/director-general/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020. Since then, the number of global cases has been exponentially growing along with the number of deaths. At the time of writing of this manuscript, the total number of worldwide cases is reported to be more than 199 million, while the total number of deaths is nearly 4 million, as per the WHO https://covid19.who.int/. Despite the tireless efforts of medical teams and researchers to put an end to this crisis, COVID-19 remains a phantom that haunts our lives to this day and research is still needed to combat this epidemic.
SARS-CoV-2 was linked from the start of the outbreak to the coronavirus family and especially to the betacoronavirus subfamily.1 The new coronavirus was speculated to be linked in some manner to HIV-1 and the bat SARS-like coronavirus. It is, however, confirmed that SARS-CoV-2 is derived from SARS-CoV with a 79% sequence identity between both viruses.2-4 The virus SARS-CoV-2 genome is comprised of 29.9 kb arranged in a single strand positive sense RNA molecule.5 The structure of SARS-CoV-2 consists of 4 structural proteins and 16 non-structural proteins (NSPs). The most important COVID-19 therapeutic targets are Nsp5 and Nsp12. Nsp5 is the main protease of the SARS-CoV-2 virus that processes the viral polyprotein into 16 functional proteins.6 Nsp12 is the viral RNA-dependent RNA polymerase (RdRp) which participates in viral replication.7
The RdRp enzyme is a key regulator of viral replication and transcription of the viral genome.8 This enzyme synthesizes an RNA strand from a template RNA strand. The RdRp activity of Nsp12 is largely compromised in the absence of Nsp7 and Nsp8. The presence of both non-structural proteins as co-factors to the RdRp along with the template RNA is essential for its activity. The RdRp contains a nidovirus RdRp-associated nucleotidyltransferase (NiRAN) structure at the N-terminal. The C-terminal contains an RdRp domain with a right-hand configuration and three subdomains; the palm, the fingers and the thumb subdomains. A unique β-hairpin stabilizes the overall structure of Nsp12 by forming interactions with the NiRAN domain and the palm subdomain. The palm subdomain contains the active site, where the two most important residues are Asp760 and Asp761.
A number of inhibitors have been reported for SARS-CoV-2 RdRp. The reports include compounds that act as nucleoside analogues as well as non-nucleoside analogue RdRp inhibitors. Discovery approaches of RdRp inhibitors include computational screening approaches, lab testing and drug repurposing approaches.9,10 As a result of this work, remdesivir was approved for use in severe COVID-19 cases.11 This step has encouraged the investigation of new compounds for their potential use as RdRp inhibitors to be clinically beneficial for COVID-19 patients. In this work, a natural product database composed of nearly 8580 compounds is investigated through a rigorous computational approach. The workflow aimed to filtering the database through a docking screen of increasing precision. Molecular docking is a computational experiment that aims to identify possible poses and interactions of a certain ligand within the binding site of biomolecule. Low precision docking screens are first used to filter large databases where the hits are ultimately filtered through docking workflows of increasing precision. The final poses and interactions of the successful ligands are then studied via inspection of the results of a highly precise docking experiment called extra precision docking (XP).12 The filtered compounds were then investigated for additional properties and pharmacophoric features that underscore their binding to RdRp. Through this work, two potential RdRp natural product inhibitors were identified; Sennoside B and Ginsenoside B2.
Ligand and protein preparation
A total of 8576 natural products ligand structures were obtained from the Zinc Database.13 All compounds were obtained from natural compound databases (AfroDB Natural Products,14 TimTec Natural Derivatives http://www.timtec.net, Specs Natural Products https://www.specs.net/index.php, Indofine Natural Products http://www.indofinechemical.com/Media/sdf/sdf_files.aspx and UEFS Natural Products zinc.docking.org/catalogs/uefsnp available through the Zinc Database. All databases were accessed on 06/04/2021. The downloaded ligands were prepared using Ligprep and the used force field was OPLS3 Schrödinger Release 2021-2: LigPrep, Schrödinger, LLC, New York, NY, 2021. The crystal structure of the SARS-CoV-2 RdRp was downloaded from the Protein Data Bank (PDB).15 The downloaded crystal structure contained the RdRp itself co-crystallized with the viral protein cofactors Nsp7 and Nsp8 (PDB ID: 6M71).16 The downloaded protein was then prepared using the Protein Preparation Wizard and standard protein preparation criteria.17
Docking grid generation and sequential docking protocol
To conduct the ligand docking experiment, the prepared crystal structure of the RdRp was used to generate a docking grid. The docking grid was generated using the Grid Generation protocol.18 The generated grid box had central coordinates 112.43, 114.18 and 130.26 for x, y and z dimensions respectively. No constraints were generated nor any excluded volumes or rotatable groups to fully explore the best binding pose for each ligand within the vicinity of the enzyme.
The docking experiment was done in three different steps, all using Glide.18 The first docking step was performed using high throughput virtual screening (HTVS) where 8576 ligands were docked to produce 8372 poses.19 The following docking step was performed on 840 ligands to produce 840 poses using standard precision (SP) docking.18 The final docking step was performed on 22 ligands to produce 22 poses using extra precision docking (XP).20 The poses were then inspected manually using the 2D and 3D pose viewers in Maestro Schrödinger Release 2021-2: Maestro, Schrödinger, LLC, New York, NY, 2021.
Pharmacophore generation
The 22 ligands used for XP docking were used to generate a pharmacophore hypothesis using Phase.21 An XP docking score threshold was used to designate active and inactive compounds for the pharmacophore model to be generated. The threshold was set to be -7.000 where compounds having higher score were assigned to be active and those with lower score were assigned to be inactive. The threshold of -7.000 was chosen since compounds of lower docking score showed visible clashes and/or very few interactions with active site residues upon visual inspection of their poses. 12 compounds were therefore assigned as active while the remaining 10 compounds were inactive based on this criterion. The produced pharmacophore was used in another experiment to screen the predicted compound metabolites also using Phase.
Metabolite and ADME properties prediction
The two compounds of highest score from the XP docking experiment were evaluated for their potential metabolites. Metabolite prediction was done using GLORY to predict the potential metabolites resulting from Cyp450 processing, while GLORYx was used to predict phase I and II metabolites.22,23 ADME properties were studied using QikProp Schrödinger Release 2021-2: QikProp, Schrödinger, LLC, New York, NY, 2021. The studied properties included molecular weight calculation, solvent accessible surface area (SASA) and solvent accessible volume.
Results of natural products docking screen against SARS-CoV-2 RdRp
In this work, a total of 8576 entry compounds of natural sources were docked into the RNA-binding tunnel of the SARS-CoV-2 RdRp. A high-throughput virtual screening (HTVS) approach was utilized to filter potentially useful compounds and identify hit compounds. The docked compounds resulted in 8372 poses after excluding duplicates and ligands with excessively large size. Excluded ligands included those with more than 500 atoms, more than 100 rotatable bonds and/or length more than 20 Å. The resulted docking poses showed docking score ranging from -7.462 to 2.452 kcal/mol. Positive docking scores indicate non-spontaneous binding while negative scores indicate spontaneous favorable binding down the energy gradient. Out of all the resultant docking poses, only 29 ligands showed positive docking scores.
The HTVS step was only a preliminary screening to filter out compounds of no practical affinity to the SARS-CoV-2 RdRp. A standard precision docking experiment was then conducted involving the top 840 compounds from the HTVS screen (~top 10% of the total docked ligands). This experiment aimed to further refine the molecular docking and pick out ligands of superior activity. The docked ligands had an HTVS docking score ranging from -7.462 to -4.469 kcal/mol while their more accurate standard precision docking scores ranged from -8.226 to -2.591kcal/mol. About 516 of the docked compounds showed scores higher than remdesivir triphosphate which had a standard precision docking score of -4.999kcal/mol. The average molecular weight of the compounds displaying higher SP docking score than remdesivir triphosphate was 463.5 amu which is lower than remdesivir triphosphate which is 531.2 amu. However, the average solvent-accessible surface area (SASA) of those compounds was 715.1 Å2, only slightly lower than remdesivir triphosphate which has a SASA of 748.4 Å2. Furthermore, the average total solvent-accessible volume of the compounds was 1320.6 Å3, slightly higher than that of remdesivir triphosphate which is 1315.3 Å3.
Extra precision docking of selected compounds against SARS-CoV-2 RdRp
Docking results of the SP docking run showed that the compound database included a number of compounds with potential activity against SARS-CoV-2 RdRp. The top scoring compounds were selected for further extra precision (XP) docking to accurately predict their binding poses and affinities. The selected compounds were the ones having a docking score of -7.000kcal/mol or more. 22 compounds matching with this criterion were selected. The XP docking scores of these compounds ranged from -11.534 to -2.692 kcal/mol. 4 of the docked compounds had an XP docking score higher than remdesivir triphosphate, whose XP docking score was -8.696 (Table 1). The average molecular weight of those 4 compounds is 877.4 amu, much higher than remdesivir triphosphate. The average values of SASA and solvent-accessible volume of the 4 compounds are 1156.4 Å2 and 2557.6 Å3 respectively, much greater than the values of remdesivir.
Table 1 Compounds having XP score higher than remdesivir
Pharmacophore model prediction for natural SARS-CoV-2 RdRp inhibitors
To find a relationship between the top scoring compounds, a pharmacophore model was generated using the top 22 compounds previously docked with XP settings. 12 of the 22 compounds were assigned as “active” compounds while the remaining 10 compounds were assigned as “inactive”. This assignment was based on a docking score threshold from the XP docking experiment to define common pharmacophoric features of the highest-ranking compounds while excluding features of compounds with lower docking scores. The best generated pharmacophore hypothesis matched 8 of the “active” compounds. This hypothesis includes a total of 5 features; 2 hydrophobic features, 1 aromatic ring feature, 1 hydrogen bond donor and 1 hydrogen bond acceptor (Figure 1).
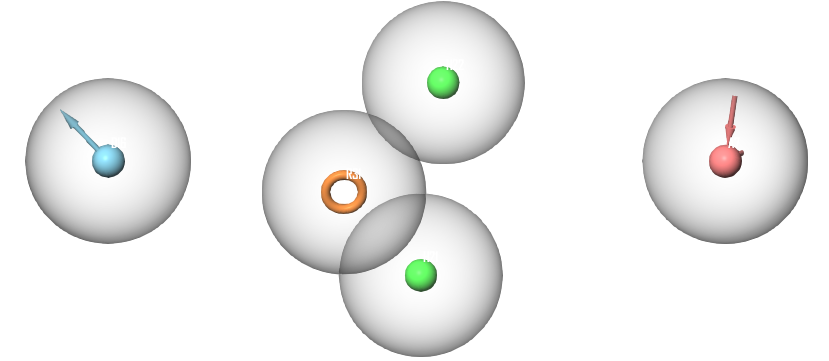
Figure 1 Pharmacophoric features of the highest scoring compounds. The displayed features include 2 hydrophobic features (green), one aromatic ring feature (orange), one hydrogen bond donor feature (blue) and one hydrogen bond acceptor feature (red).
Metabolite prediction and assessment of metabolite activity
Since the activity of any compound may also be attributed to the activity of one or more of its metabolites, a metabolite prediction experiment was conducted. The metabolites of the two most active and commercially available compound, Sennoside B and Ginsenoside B2, were predicted using GLORY and GLORYx.22,23 Possible metabolites of phase I and II and CYP-450 metabolites were generated and the most likely metabolites were picked (Table 2). Sennoside B showed 2 possible metabolites, one resulting from the cleavage of one of the sugar rings (MSB1), while the other resulted from the complete removal of one of the sugars (MSB2). Another important metabolite of Sennoside B, Rhein Anthrone (MSB3), was also added to the series as it is a known metabolic biproduct through gut bacteria.24 Ginsenoside B2 showed only one notable metabolite, a hydroxylated compound at the allylic carbon (MGB1).
|
Compound ID |
Compound Structure |
XP Docking Score |
Pharmacophore Fit Value |
|
MSB1 |
 |
-12.306 |
1.522 |
|
MSB2 |
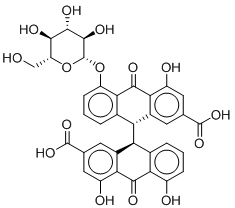 |
-8.929 |
1.517 |
|
MSB3 |
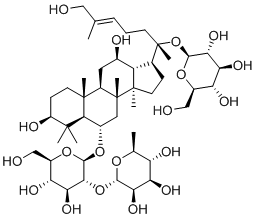 |
-4.877 |
1.124 |
|
MGB1 |
-9.939 |
1.516 |
Table 2 Metabolites of Sennoside and Ginsenoside B2
To determine the possible anti-SARS-CoV-2 RdRp activity of the predicted metabolites, and XP docking experiment was performed in a manner similar to that conducted on the original ligands. Three of the four tested metabolites showed XP docking scores higher than remdesivir triphosphate. MSB1 showed a docking score superior to Sennoside B while MSB2 scored lower than Sennoside B but higher than remdesivir triphosphate. MSB3 had a very low docking score compared to both Sennoside B and remdesivir triphosphate. MGB1 had a score higher than remdesivir but lower than Ginsenoside B2.
To further assess the fitness of the metabolites to the pharmacophore hypothesis of SARS-CoV-2 RdRp inhibitors, the compounds were screened against the generated pharmacophore. MSB1 and MSB2 displayed a fitness score of 1.522 and 1.517 respectively. Both compounds had a fitness score higher than Sennoside B whose fitness score was 1.248. MSB3, however, had a fitness score lower than Sennoside B, scoring only 1.129. On the other hand, MGB1 scored slightly lower than Ginsenoside B, scoring 1.516 while Ginsenoside B scored 1.552. These pharmacophore fitness results are in accordance with the docking results of the metabolites.
The search for natural products with anti-RdRp activity started by screening a database of 8576 natural product compounds obtained from variable sources through the Zinc Database. A high throughput virtual screen protocol showed that most of the compounds included in the initial screen possessed positive binding affinity to the RdrP RNA binding tunnel. The top 10% of the docked compounds were chosen based on HTVS docking scores to be further evaluated through a more precise docking experiment. Standard precision docking further segregated these compounds based on binding affinity in terms of docking scores. Further filtration of the docked compounds based on SP docking scores produced 22 final compounds for more in-depth assessment through XP docking.
The top two compounds from the screening protocol showed interesting binding modes that somewhat compared to that of remdesivir triphosphate but warranted superior binding affinity. Remdesivir is a phosphoramidate prodrug metabolized by cells to produce the active form, remdesivir triphosphate.25 Once activated, remdesivir triphosphate binds to the tip of the nucleoside triphosphate (NTP) channel in the RdRp complex with elongating RNA. Computational, biochemical and synthetic RNA therapeutic approaches showed that remdesivir triphosphate hinders the elongation of the RNA chain by preventing the incorporation of nucleotides after 3 new nucleotides are inserted into the growing chain following the drug itself.26,27
Sennoside B was found to bind to the NTP channel but not only at the tip, it spanned the entire channel and made interactions with active site residues. The symmetric structure of sennoside B allowed the placement of the central dianthrone nucleus at the core of the RdRp channel. One of the anthrone rings made hydrogen bonding interactions with the backbone carbonyl of Ser759 and the carboxylic acid side chain of Asp761. It is important to note that Asp761 is a key active site residue involved in the catalytic reaction of RdRp.28 The sugar groups interacted with the termini of the RdRp channel to further anchor the compound within the channel. The first sugar group formed three hydrogen bonds, two of them with the carboxylic acid side chain of Asp623 and one with the amide side chain of Asn691. The other sugar group also forms three hydrogen bonds, two with the backbone carbonyl of Thr591 and one with the nitrogen of Gly590. The almost semi-helical structure of Sennoside B allows it to spiral through the channel of RdRp, in a manner similar to that of RNA. This allows for the smooth and strong binding without any clashes between Sennoside B and the binding site (Figure 2).
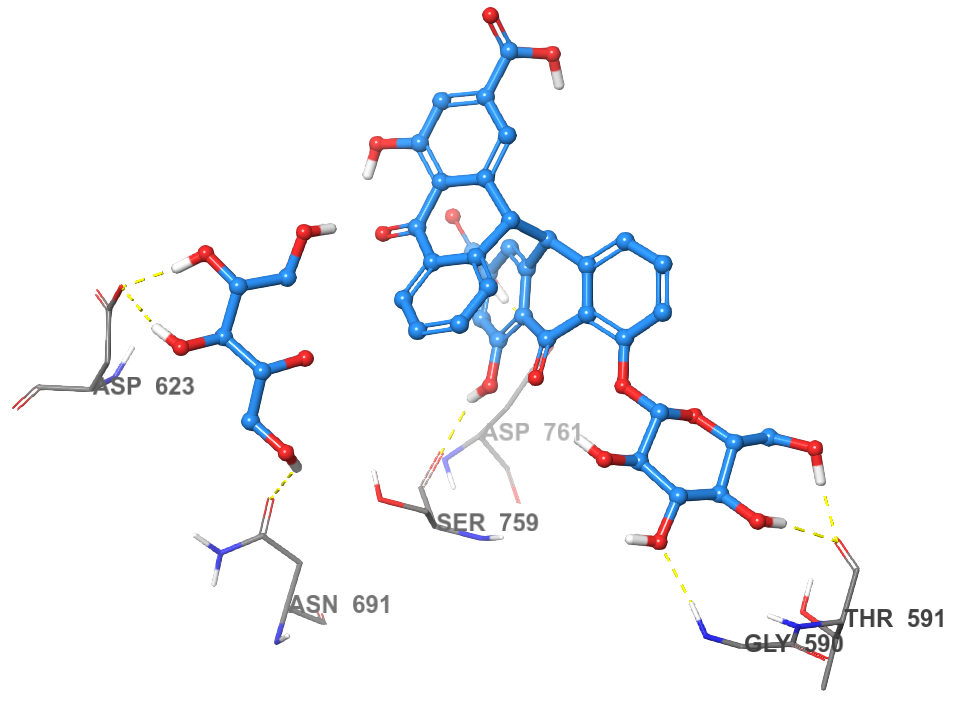
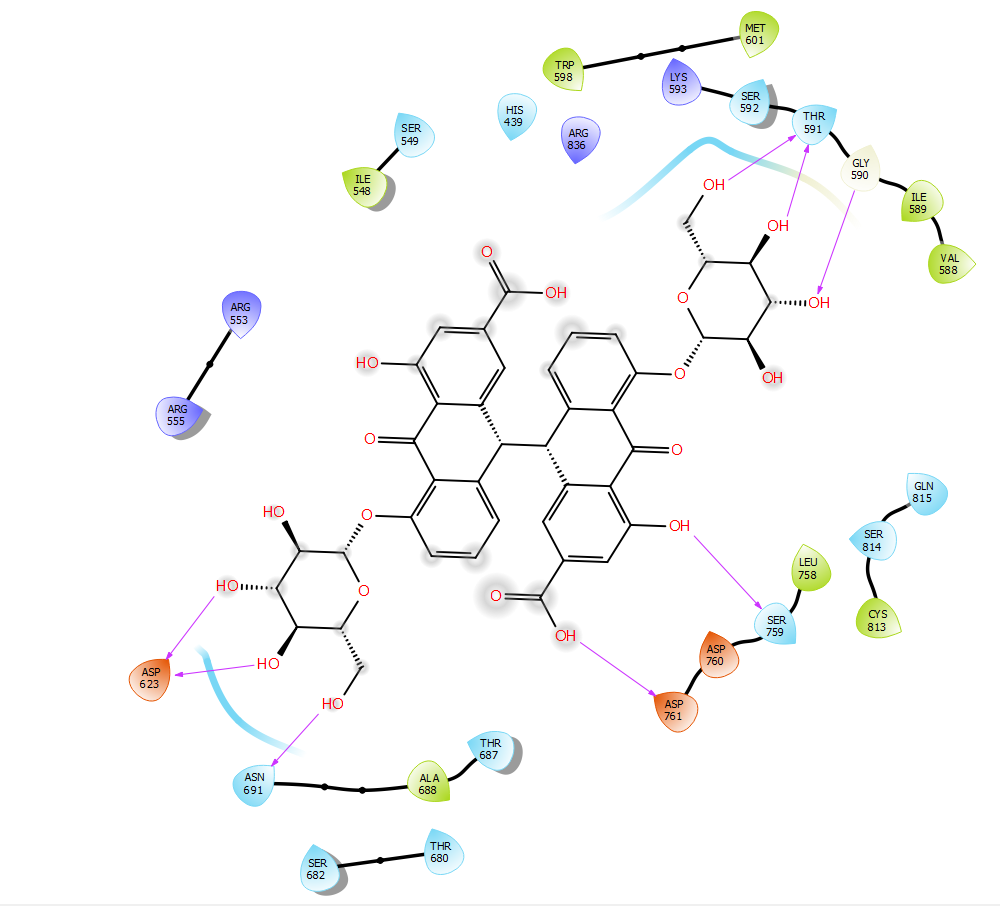
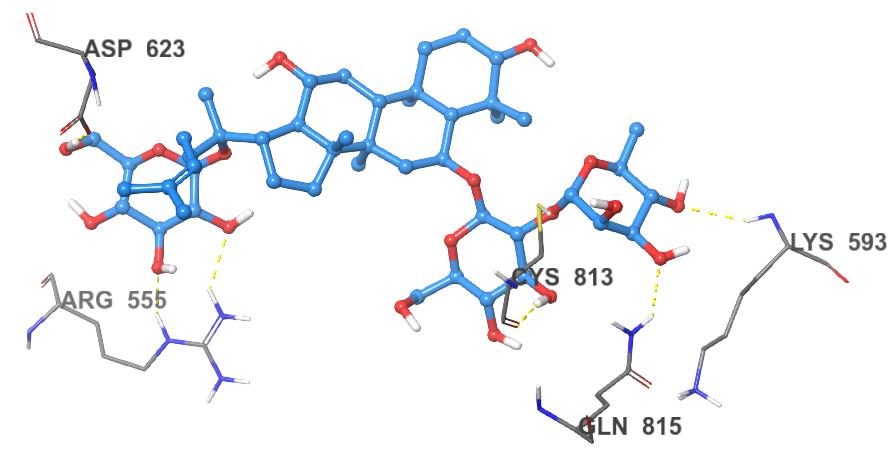
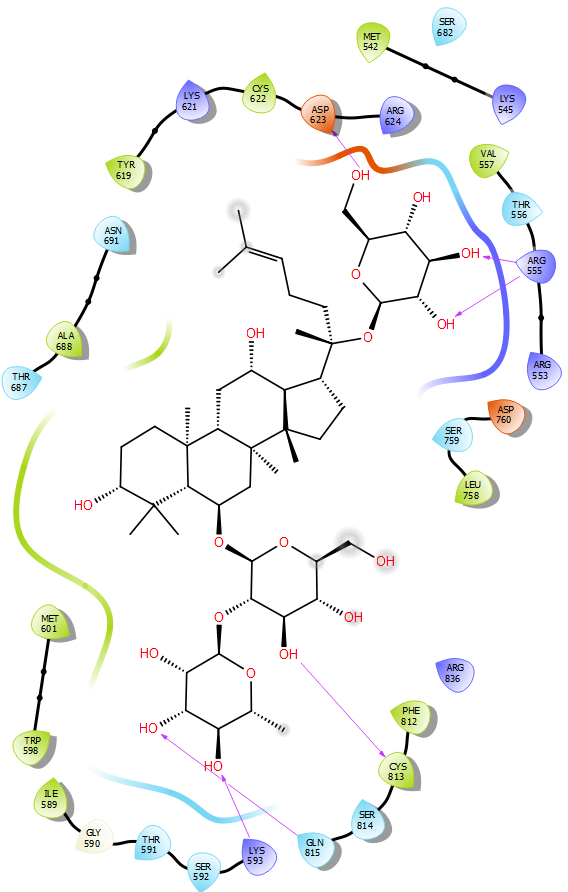
Figure 2 The 2D and 3D binding poses of Sennoside B (top) and the 2D and 3D binding poses of Ginsenoside B2 (bottom).
Compared to Sennoside B, the hydrolysis metabolic product MSB1 shows much more binding interactions with RdRp. It is able to form 11 hydrogen bonding interactions with 7 different amino acids in the vicinity of the binding pocket. The amino acids hydrogen bonding with MSB1 are Ser549, Lys551, Arg553, Tyr619, Asp623, Asp760 and Arg836. Additionally, one of the anthrone rings was able to form a π-cation interaction with the side chain of Arg555. Although this compound was not able to interact with one of the catalytic residues, it was able to form extensive non-bonding interactions at the entry of the NTP channel. The compound is able to twist its confirmation to completely block the entry of the channel, thereby possibly preventing the entry of a new NTP to join the elongating RNA.
The other metabolite of Sennoside B that shows a high docking score, MSB2, is shown in its docking pose to lodge itself sideways at the entrance of the NTP channel. This compound is not as large as Sennoside B since it completely lacks one of the two sugar moieties, therefore it is not able to extend all the way across the RdRp channel. Despite this difference in binding mode, MSB2 is also still able to completely block the NTP channel. A total of 7 hydrogen bonds are formed between MSB2 and 7 different amino acids in the RdRp channel. The hydrogen bonding amino acids are Arg553, Lys621, Asp623, Asn691, Lys798, Trp800 and Glu811. Also, similar to the MSB1, MSB2 forms a π-cation interaction but with the side chain of Arg553 instead of Arg555. Unlike MSB1 and MSB2, MSB3 forms very few interactions with RdRp channel residues. The small metabolite binds against the side of the RdRp channel within its vicinity, away from the entrance and exit of the tunnel. It is able to form only 4 hydrogen bonds with 3 residues, Arg553, Arg555 and Thr556. The loose binding and insignificant binding site of the compound makes it an unlikely inhibitor of the enzyme.
Ginsenoside B2 is found to bind to the RdRp channel in a manner similar to that of Sennoside B. The large structure of the compound allows it to span across the length of the RdRp channel, anchoring itself to the entry and exit. A network of hydrogen bonds between the compound’s sugar moieties and residues at the entrance and exit of the tunnel. The first sugar moiety forms two hydrogen bonds with the side chain of Arg555 and one hydrogen bond with the side chain of Asp623. The second sugar moiety forms one hydrogen bond with the amide nitrogen of Lys593, a second hydrogen bond with the carbonyl of Cys813 and a final hydrogen bond with the side chain of Gln815. However, Ginsenoside B2 does not form any interactions with catalytic residues of the enzyme. Furthermore, some clashes are observed between the compound and the side chains of Arg555 and Arg553. For those reasons, Ginsenoside B2 is found to have lower affinity to RdRp than Sennoside B (Figure 2).
The metabolite of Ginsenoside B2, MGB1, is found to possess an elevated docking score and a possibly high affinity to RdRp. The metabolite readjusts its binding pose in comparison to the original compound where it is lodged at the entrance of the NTP channel in a sideways manner completely blocking the entrance. The compound forms 7 hydrogen bonds to 6 different amin acids; Lys545, Arg553, Thr556, Tyr619, Asn623 and Asp760.
A very important finding is that most of the high-scoring compounds in all three docking screens (HTVS, SP and XP) possessed a relatively large molecular weight, SASA and solvent-accessible volume. This finding is in agreement with the observed binding mode of the screened compounds. Large compounds and metabolites are able to lodge themselves either at the entrance of the NTP channel or across the entire channel. The small metabolite MSB3 displays a low docking score and an insignificant binding mode that makes this compound an unlikely inhibitor of RdRp. These results show that the large size of the inhibitor plays an important role in its binding affinity.
The pharmacophoric features of the inhibitors clearly correlate the structures of the inhibitors. The two most important features are the hydrogen bond donor and acceptor with a distance of 14.5 Å. These features warrant the hydrogen bonding of the inhibitor termini to the entrance and exit of the tunnel or to the residues across the tunnel to anchor the inhibitor sideways. The aromatic feature, although not essential in the binding poses of all the compounds, gives the probability for extra π-cation interaction with the side chains of Arg553 or Ag555. The hydrophobic pharmacophoric features represent the core of the compound to maintain the proper distance between the hydrogen bond donor/acceptor features.
Although Sennoside B and Ginsenoside B2 are very large natural products that violate the Lipinski rule of 5 and if administered orally only act locally in the gastrointestinal tract, the possibility of their intravenous administration can be investigated. Recently, a study by Abdallah et al. described a potent inhibitory activity of Sennoside B against SARS-CoV-2 main protease where the IC50 of the compound was found to be 104nM.29 Combined with the results in this study, Sennoside B may be able to act as a dual inhibitor of both SARS-CoV-2 main protease and RdRp.
In this project, a computational biochemical workflow was constructed to discover and repurpose commercially available natural products as SARS-CoV-2 RdRp inhibitors for potential therapeutic purposes. The workflow started by screening a library of ~8580 natural compounds obtained from the Zinc Database, followed by filtration of the compounds based on docking scores through SP and XP docking. Along with the docking study, an investigation of some molecular properties such as molecular weight, SASA and solvent-accessible volume were investigated to better understand the relation between such molecular properties and the binding mechanism of the compounds. A pharmacophore hypothesis was constructed to study the structural determinants of the inhibitors. Two candidate compounds were further investigated through metabolite prediction and XP molecular docking of the most probable metabolites. This study sheds light onto large molecular weight natural products as potential therapeutic non-covalent inhibitors of SARS-CoV-2 RdRp. Sennoside B and Ginsenoside B2 are candidate natural products that may play an important role in RdRp inhibition. We encourage other research groups to conduct further in vitro and in vivo experiments to confirm these in silico results and establish a new role for previously known natural compounds.
None.
Author declares that there is no conflict of interest.

©2021 Sharaf. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.