Journal of
eISSN: 2377-4282


Research Article Volume 7 Issue 2
College of Medicine, University of Al-Qadisiyah, Iraq
Correspondence: Hasan R Khudhur, College of Medicine, University of Al-Qadisiyah, Iraq,, Tel 009647808360603
Received: February 07, 2017 | Published: April 17, 2018
Citation: Alomashi GB, Khudhur HR. MIF G-173C polymorphism and susceptibility to cutaneous leishmaniasis in Iraq. J Nanomed Res. 2018;7(2):89 ? 91. DOI: 10.15406/jnmr.2018.07.00183
Background/purpose: Cutaneous leishmaniasis (CL) is a parasitic disease transmitted by biting of the sandfly; it is a severe health problem in many regions of the world. The disease is endemic in Iraq and other countries. In Iraq, there are two main species of the genus Leishmania causing the infection: L. tropica and L. major. Previous studies refer to that a genetic makeup of host also have an essential role in the outcome of the disease, the present study investigates the association between Cutaneous Leishmaniasis and functionally active polymorphisms in the Macrophage migration inhibitory factor (MIF gene).
Methods: Samples of peripheral blood were collected from 60 patients with CL and 32 apparently healthy controls. MIF G-173C polymorphism was detected in patients and control groups by PCR-RFLP technique.
Results: There was no statistically significant difference in the MIF G-173C polymorphism between the CL patients and healthy controls (P=0.234), as well as no association between MIF G-173C polymorphisms and cutaneous leishmaniasis, were the frequency of allele C showed a trend towards less frequency seen in patients compression to that observed in controls group (P = 0.512).
Conclusion: MIF G-173C polymorphisms do not affect susceptibility to cutaneous leishmaniasis.
Keywords: cutaneous leishmaniasis, genetic susceptibility, MIF G-173C polymorphisms
Human leishmaniasis is a parasitic disease transmitted by sand flies, its characteristic by a spectrum of cutaneous, mucocutaneous and visceral diseases that depend largely on the species of the parasite involved and host immune response.1,2 Cutaneous leishmaniasis is the most common form of leishmaniasis, about (1-1.5) million of cases every year, and about (50 to 70%) of all cases in the world.2,3 Cutaneous leishmaniasis occurs each year more than 90% of cases occur in five countries in the old word (Afghanistan, Algeria, Iran, Iraq and Saudi Arabia) and two countries in the new world including Brazil and Peru.4 Leishmania major and Leishmania tropica considered as common causes of Cutaneous leishmaniasis in Iraq.5 Macrophage migration inhibitory factor (MIF) is considered to be one of the first cytokines to be discovered, its consider an essential component of the immune response of host against microbial and induce activation and secretion of interleukins like TNF-α, IFN-γ, IL -1β, IL-12, IL-6 and IL-8 by immune cells.6 MIF increase survival of macrophage by inhibition activity of P53 and thus decrease activation-induced apoptosis.7,8 Finally, cDNA was cloned in 1989 in human, MIF genomic localization to chromosome 22q11 later mapped, the human MIF gene has three exons of 205, 173 and 183 bp, these are separated by two introns of 189 and 95 bp.6,9 Previous study refer to that MIF plays an essential role in resistance of host to Cutaneous leishmaniasis, were found human MIF activate infected macrophage to kill L. major at a concentration (1.5 -2.5μg/ ml) in vitro.10 Jesus et al in Brazil found that associated between MIF-173 C polymorphism and cutaneous leishmaniasis.11 Jesus suggest that the MIF-173C allele induce lower levels of MIF cytokine in serum, and this lower synthesis of MIF might behave correlation with susceptibility to leishmaniasis. The present study aims to investigate of MIF-173 C polymorphism with susceptibility to CL infection in Iraqi population in AL-Muthanna province.
Subjects and study design
A total of 60 patients infected with CL selected from areas endemic for Cutaneous leishmaniasis (CL, 34 male, and 26 female, mean age = 17±13years old), Patients were clinically diagnosed as patients with cutaneous leishmaniasis based on Dermatologist diagnosis as well as clinical and parasitological parameters.12 While negative control group (18 males and 14 females; mean age = 21±9 years old), those included 32 apparently healthy people without any cutaneous lesions and not suffer from any disease.
MIF -173G/C typing
Genomic DNA from blood samples was extracted by using Accupower® Genomic DNA extraction kit (Whole Blood), the extracted genomic DNA from blood samples was checked by using Nanodrop spectrophotometer, that check and measurement the purity of DNA through reading the absorbance in at (260 /280 nm). Genotyping for MIF -173G/C polymorphisms was performed by PCR-RFLP with two primers as described by13 and provided from Bioner Company were listed in the Table 1. The volume of PCR Mixture for polymorphisms was 50 μl, including Pre master mix (Bioneer, Korea), 5 μl (20 ng/mL) of genomic DNA, 2.5mL of each primer reverse and forward (10pm/ml) and volume complete with Nuclease-free water. Thermocycler conditions were: 94°C for 5 minutes then 38 cycles of 94°C for 30 second, 59°C for 30 seconds, and 72°C for 30 second followed by 5-minute extension at 72°C. Then PCR products were analyzed by agarose gel electrophoresis on a 1% agarose gel and stained with ethidium bromide (10 mg/ml). The PCR products digested by using restriction enzyme AluI, and RFLP- PCR master mix did according to company instructions (Biolabs/ U.K). After that, RFLP-PCR product was analyzed by agarose gel electrophoresis (2.5%) and product analysis
Name |
Primer sequence |
R. enzyme |
Genotype |
Products |
MIF173G/C |
F:5’-ACT-AAGAAA-GAC-CCG-AGG-C-3 |
AluI |
GG |
268 and 98bp |
R:5’-GGG-GCA-CGT-TGG-TGTTTA-C-3 |
CC |
268,205.98 and 63bp |
||
CG |
205,98 and 63bp |
Table 1 The sequence of primers, restriction enzyme, and DNA Fragments size
Statistical analysis
Statistical analysis was conducted by using (SPSS 23). Determine the statistical differences among different groups and associations between allelic and genotypes of MIF -173G/C polymorphisms with susceptibility to CL was performed by using the Pearson Chi-square (ᵡ2) test and odds ratio with (95%) confidence interval.14 The probability of (P≤ 0.05) was considered to be statistically significant.
In total, 60 CL patients and 32 healthy controls were genotyped, equilibrium of Hardy-Weinberg was performed in both patients and controls. Figure 1 & Table 2 shows Distribution of MIF -173 genotypes and alleles among patients and controls, were detected by PCR-RFLP technique, at this locus there is three genotypes; GG, GC, and CC with band sizes 268/98 pb, 205/ 268/98/63 pb and 205/98 pb respectively. A no significant difference in genotype distribution between CL patients and healthy controls (P=0.234). These study refer towards the high frequency of GG genotype among patients (44/60, 73.30%) when compared with healthy controls (20/32, 62.5%) were (P=0.282). Moreover there are no statistical significant (P=0.512) in G and C allele between patients and healthy controls, it was found that high frequency of G allele in patients groups 102(85%) compression with G allele 52(81.25%) in the control group (P=0.512), and less frequency of C allele in patients groups 18(15%) compression with observed in G allele 12(18.75) in the control group (P=0.512).
Variable |
Patients(60) |
Control(32) |
P value |
Odd ratio |
95 % Confidence |
|
|
|
|
|
|
Lower |
Upper |
|
|
|
|
|
|
|
GG |
44(73.30%) |
20(62.5%) |
P=0.282 |
1.650 |
0.660 |
4.125 |
GC |
14 (23.3%) |
12(37.5%) |
P=0.151 |
0.507 |
0.200 |
1.289 |
CC |
2 (3.3%) |
0(0%) |
- |
- |
- |
- |
Allele |
|
|
|
|
|
|
G |
102(85%) |
52(81.25%) |
P=0.512 |
1.308 |
0.586 |
2.920 |
C |
18 (15%) |
12(18.75) |
P=0.512 |
0.765 |
0.343 |
1.707 |
Table 2 Distribution of MIF -173 genotypes and alleles
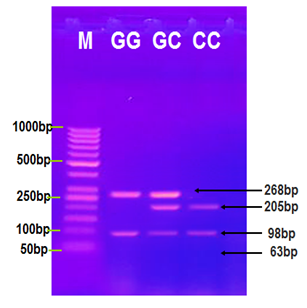
Figure 1 Agarose gel electrophoresis image that shows the RFLP-PCR product analysis of MIF gene by using AluI restriction enzyme. Where M: marker (1000-50bp), lane (GG) homozygote product digested with a restriction enzyme at 268bp and 98 bp bands. Lane (G/C) heterozygote, the product digested by restriction enzyme into 268bp, 205bp, 98bp and invisible 63bp bands. Lane (CC) homozygote, the product digested by restriction enzyme into 205bp, 98bp and invisible 63bp bands.
This study investigates the association between Cutaneous Leishmaniasis and functionally active polymorphisms in the MIF gene. The result of Detection of MIF -173 polymorphism among patients and controls were agreement with the previous study.15 In MIF -173 polymorphism, were no significant difference in genotype distribution between Cutaneous Leishmaniasis patients and healthy controls (P=0.234), and no significantly (P=0.512) in G and C allele between patients and healthy controls. However, current result refers to that no association between MIF G-173C polymorphisms and cutaneous leishmaniasis, which contrast with the previous study confirmed in Brazil which refers to finding a correlation between MIF-173 C polymorphism and cutaneous leishmaniasis.11 However, other factors should be taken into accounts, such as variant in the genetic makeup of their populations, or possible another factor like the (MIF-794CATT5-8) polymorphism and might be a correlation with different infecting Leishmania species, animal host species and vectors found in this area.16 Therefore, all these factors may be plays essential role in susceptibility to CL.
None.
The authors declare that no competing interest exists.

©2018 Alomashi, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.