The Pranav distribution is defined by its probability density function (pdf) and the Cumulative density function (cdf)
(1.1)
( 1.2)
It should be noted that the Pranav distribution, a convex combination of exponential
and gamma
distributions, has been proposed by Shukla1 for modeling lifetime data. Important statistical properties of Pranav distribution including its shapes, moments, skewness, kurtosis, hazard rate function, mean residual life function, stochastic ordering, mean deviations, Bonferroni and Lorenz curves, Renyi entropy measure and stress-strength reliability are available in Shukla.1 The Pranav distribution has been found to provide a better fit for survival time data over exponential distribution, Lindley distribution introduced by Lindley,2 Akash distribution proposed by Shanker3 and Ishita distribution suggested by Shanker and Shukla.4 The pdf of Lindley, Akash, and Ishita distributions has been presented in Table 1.
|
Lifetime distributions Pdf
|
Mixtures of distributions
|
Introducer (year)
|
|
Lindley
|
|
exponential
and gamma
distributions
|
Lindley2
|
|
Akash
|
|
exponential
and gamma
distributions
|
Shanker3
|
|
Ishita
|
|
Exponential
and gamma
distributions
|
Shanker & Shukla4
|
Table 1 The pdf of Lindley, Akash and Ishita distributions
Ghitany, et al5 have studied in detail on Lindley distribution. Shanker et al.6 have detailed comparative study on modeling of various lifetime data using exponential and Lindley distributions. Further, Shanker et al.7 have detailed comparative study on modeling of real lifetime data using Akash, Lindley and exponential distributions.
During recent decades several one parameter lifetime distributions have been introduced in statistics literature and the Poisson mixture of these distributions, namely Poisson-Lindley distribution (PLD) proposed by Sankaran,8 Poisson-Akash distribution (PAD) introduced by Shanker9 and Poisson-Ishita distribution (PID) suggested by Shukla & Shanker,10 are some among others.
The probability mass function (pmf) of PLD, PAD and PID has been presented in Table 2.
|
Distributions
|
Pmf
|
Mixtures of distributions
|
Introducer (year)
|
|
PLD
|
|
Poisson mixture of Lindley
|
Sankaran8
|
|
PAD
|
|
Poisson mixture of Akash
|
Shanker9
|
|
PID
|
|
Poisson mixture of Ishita
|
Shukla & Shanker10
|
Table 2 Pmfs of PLD, PAD and PID
Detailed study of PLD, PAD and PID are available in Ghitany & A1 Mutairi,11 Shanker,9 & Shukla & Shanker,10 respectively. Shanker and Hagos12 has detailed study on applications of PLD in various fields of knowledge.
The main reasons and motivation of introducing Poisson-Pranav distribution (PPD) are (i) it has been observed that Pranav distribution gives a better fit than exponential, Lindley, Akash and Ishita distributions and (ii) it is expected that PPD would prove to be a better model for over PLD, PAD and PID.
This paper has been divided into eight sections. The second section deals with the derivation of the pmf of PPD and its behaviour for varying values of parameter. The third section deals with raw moments and central moments of PPD and behaviour of mean and variance, coefficients of variation, skewness, kurtosis and index of dispersion for varying values of parameter. Increasing hazard rate and unimodality property of the PPD has been discussed in section four. The sections five and six deals with estimation of parameter using both the method of moment and maximum likelihood, and simulation study, respectively. Finally, the goodness of fit of the distribution and its comparative study along with conclusions have been presented in sections seven and eight respectively.
Poisson-Pranav distribution
Assuming that the parameter
of the Poisson distribution follows Pranav distribution, the Poisson mixture of Pranav distribution can be obtained as
(2.1)
(2.2)
This is named as Poisson-Pranav distribution (PPD)”. The pmf of PPD presented in Figure 1.
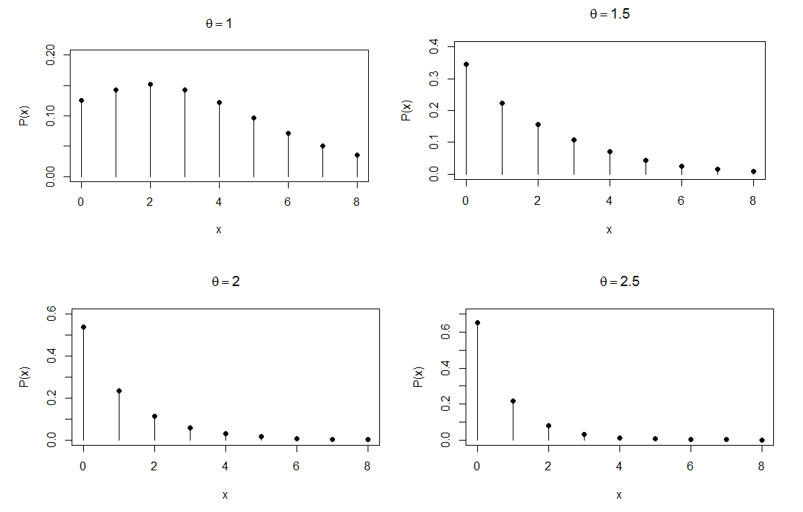
Figure 1 Behavior of the PPD for varying values of the parameter
Moments
The
th factorial moment about origin of PPD (2.2) can be obtained as
, where
Using (2.1), the
th factorial moment about origin of PPD (2.2) can be obtained as
Taking
in place of
within the bracket, we get
After simplification, the
th factorial moment about origin of PPD can be expressed as
(3.1)
Substituting
in (3.1), the first four factorial moments about origin can be obtained and using the relationship between factorial moments about origin and moments about origin, the first four moment about origin of PPD are obtained as
The relationship between moments about mean and the moments about origin of PPD gives the moments about mean as
The coefficient of variation
, coefficient of Skewness
, coefficient of Kurtosis
, and index of dispersion
of the PPD can be obtained using following formula:
,
.
The behavior of the mean and the variance of PPD have been shown in Figure 2. Clearly PPD is always over-dispersed (variance greater than the mean).
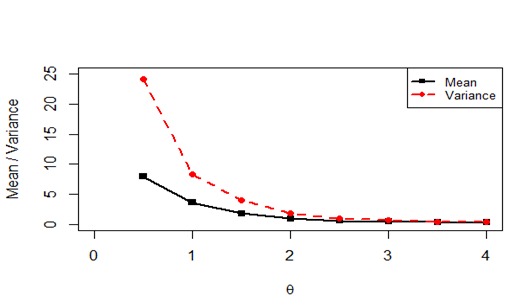
Figure 2 Plots of mean and variance for varying values of
The behavior of
of the PPD has been shown graphically for different values of parameter
in Figure 3.
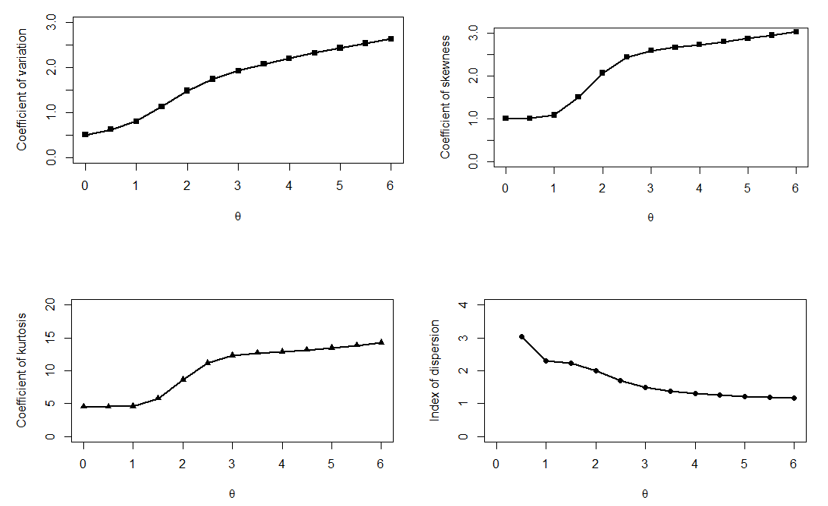
Figure 3 Behavior of coefficient of variation, coefficient of skewness, coefficient of kurtosis and Index of dispersion of PPD for different values of the parameter
Increasing hazard rate and unimodality
The PPD (2.2) has an increasing hazard rate (IHR) and thus unimodal. Clearly
is a decreasing function in
. Thus
is log-concave which means that the PPD has an increasing hazard rate (IHR) and unimodal. A detailed discussion about interrelationship between log-concavity, unimodality and IHR for discrete distributions are available in Grandell.13
Method of moment estimate (MOME)
Equating the population mean to the sample mean based on random sample
the MOME
of the parameter
of PPD is the solution of the following fifth degree polynomial equation
,
where
is the sample mean.
Maximum Likelihood Estimate (MLE)
Let
be a random sample of size
from the PPD and let
be the corresponding observed frequency The likelihood function
and the log-likelihood function of the PPD is given by
.
.
The first derivative of the log likelihood function is given by
.
The maximum likelihood estimate (MLE),
of the parameter
of PPD is the solution of the following log likelihood equation
This non-linear equation can be expressed in closed form and hence can be solved iteratively using Newton- Raphson method available in R-software. The MOME can be taken as the initial value of the parameter for Newton-Raphson method.
For a simulation study, we generate N=10,000 pseudo-random sample of sizes n=50, 100, 150, and 200 of a variable X having PPD). Then using Monte Carlo simulation we estimate the average bias and the mean squared error (MSE) of the MLEs of the parameter for
=1.5, 2, 2.5 and 3.0. The formulas for finding bias and MSE of the parameter
are
).
Using following algorithm, we generate a pseudo-random sample from PPD
Algorithm
The ML estimate, biases and the mean squares error (MSE) of the parameter based on simulated data are presented in Table 3.
|
Sample size(n)
|
|
|
|
Bias
|
MSE
|
|
50
|
1.5
|
-0.00146
|
0.0001
|
| |
2.0
|
0.00545
|
0.00148
|
| |
2.5
|
0.00237
|
0.00282
|
| |
3.0
|
0.000516
|
0.000013
|
|
100
|
1.5
|
-0.000343
|
0.000011
|
| |
2.0
|
-0.000257
|
0.000006
|
| |
2.5
|
0.001433
|
0.000205
|
| |
3.0
|
-0.00105
|
0.000111
|
|
150
|
1.5
|
-0.000433
|
0.00028
|
| |
2.0
|
-0.00002
|
0.0000006
|
| |
2.5
|
0.000123
|
0.0000022
|
| |
3.0
|
-0.00308
|
0.001431
|
|
200
|
1.5
|
0.00518
|
0.00537
|
| |
2.0
|
0.00118
|
0.00028
|
| |
2.5
|
0.00039
|
0.00003
|
| |
3.0
|
0.00088
|
0.000156
|
Table 3 Estimated Bias and MSE of MLEs
This table shows that bias and mean square error tends to zero for increasing sample size and increasing values of parameter. Further, MLE of
has a negative bias in some cases.



