Journal of
eISSN: 2469 - 2786


Review Article Volume 5 Issue 4
Departamento de Fitopatologia, Universidade Federal de Vi
Correspondence: Badel JL, Departamento de Fitopatologia, Universidade Federal de Viçosa, Avenida Peter Henry Rolfs, s/n - Campus Universitário, Viçosa, Tel +55 (31) 38992620
Received: October 03, 2017 | Published: October 18, 2017
Citation: Mendonça LBP, Zambolim L, Badel JL. Bacterial citrus diseases: major threats and recent progress. J Bacteriol Mycol Open Access. 2017;5(4):340-350. DOI: 10.15406/jbmoa.2017.05.00143
Citriculture is an important, highly organized, and competitive sector of the Brazilian economy. Nonetheless, citrus production is constantly threatened by pathogens that cause considerable economic losses and severe social impacts. Among the major citrus diseases are citrus canker (CCK), citrus variegated chlorosis (CVC), and Huanglongbing (HLB), caused by members of the bacterial species Xanthomonas citri (Xcc), Xylella fastidiosa (Xfa), and ‘Candidatus Liberibacter’ (CaL), respectively. During the lastyears, management practices for CVC and HLB in Brazil have provided good results, maintaining the disease incidence at low levels. In contrast, CCK re-emerged as the main citrus disease because of the inefficacy of the current eradication program. In this review, we discuss about the biology of the plant-pathogen interactions, several controversial aspects on the taxonomy of the causal agents, the molecular mechanisms they use to cause disease in citrus plants, the strategies used for disease management and their limitations, and some emerging control alternatives that may be available for commercial production in the future, including transgenic and genome-edited plants with enhanced resistance.
Keywords: citrus variegated chlorosis, citrus canker, huanglongbing, xylella fastidiosa, xanthomonas citri, candidatus liberibacter
CCK, citrus canker; CVC, citrus variegated chlorosis; HLB, huanglongbing; Xcc, xanthomonas citri; Xfa, xylella fastidiosa; CaL, candidatus liberibacter
Citrus is an economically important crop for at least 140 countries, which produce an annual estimate of more than 122 million tons of fruits. Brazil is the second largest citrus producer after China.1 Citrus production is a major agribusiness in Brazil, not only because of its financial volume, but also because it is a highly organized and competitive sector.2 Bacterial diseases pose a constant threat to citrus cultivation and cause substantial economic impacts in all growing areas around the world. Among them, citrus canker (CCK), citrus variegated chlorosis (CVC), and Huanglongbing (HLB) cause significant reductions in production.3-6 Between years 2000 and 2010, in the major Brazilian production area, comprising the State of Sao Paulo and the Triangulo Mineiro, these three diseases, along with citrus sudden death, were responsible for the eradication of approximately 39 million trees and losses of approximately 80 million boxes of orange per year.3 These three diseases also make it unfeasible growing citrus in some contaminated areas.3-6
In this review, we discuss the research progress on taxonomy, pathogenicity mechanisms, management, and chemical control of the causal agents of the three most economically important bacterial diseases of citrus: CCK, CVC and HLB. Although information has been published on the interaction of CCK and HLB with their insect vectors, which is relevant for disease management, we restrict our discussion to the plant-pathogen interactions.
CCK and CVC are caused by Xanthomonas citri (Xcc; sin. X. axonopodis pv. citri and X. campestris pv. citri) and Xylella fastidiosa (Xfa), respectively. The two species are phylogenetically closely related, both belonging to the family Xanthomonadaceae of the Gammaproteobateria.5,7 HLB is caused by species of the provisional genus ‘Candidatus Liberibacter’ (CaL), circumscribed in the Rhizobiacae of the Alphaproteobacteria.6,8 Three CaL species have been reported to cause HLB in citrus: ‘Ca. Liberibacter asiaticus’ (CaLas),8,9 ‘Ca. Liberibacter africanus’ (CaLaf),8,9 and ‘Ca. Liberibacter americanus’ (CaLam).10
Symptoms of CCK (Figure 1), CVC, and HLB, strategies used for their management and their current situation in Brazil are summarized in Table 1.5,6,11-20 The management of these three diseases have several aspects in common: (i) orchards must be maintained in good nutritional conditions; (ii) periodic inspections must be carried out by trained personnel; (iii) pathogen-free propagating plant material should be used;5,6,12,14,15 (iv) wind breaks should be planted; and (v) infected trees should be eradicated, except for trees older than four years infected with Xfa.12-14 Insects should be controlled in the case of CVC and HLB. In the management of CaL spp., in addition to diseased citrus trees, plants of the genus Murraya, host of both the insect vector and CaL, should also be eradicated.15 CVC is the only disease for which pruning is recommended; drastic pruning of branches in trees older than four years provides good control efficiency.17,18
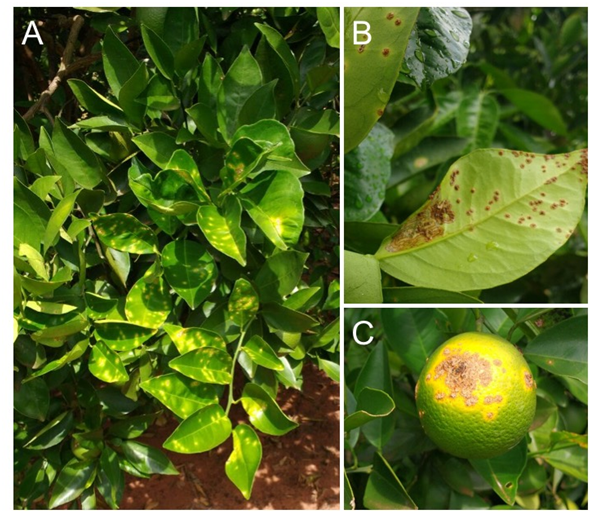

Figure 1 Symptoms caused by Xanthomonas citri in citrus plants. A - Corky spots surrounded by chlorotic haloes on the adaxial side of leaves. B - Eruptive, corky and pointed lesions surrounded by chlorotic haloes on the abaxial side of leaves. C - Larger corky lesions with centered cracks on a fruit (photos courtesy of Fabrício Eustáquio Lanza, 2017).
Disease |
Pathogen |
Symptoms |
Management |
Current Situation* |
Citrus canker (CCK) |
Xanthomonas citri |
|
|
Epidemic |
Citrus variegated chlorosis (CVC) |
Xylella fastidiosa |
|
|
Almost extinct |
Huanglongbing (HLB) |
‘Candidatus Liberibacter’ |
|
|
In check |
In contrast to the disappointing scenario currently observed for CCK for which the incidence has increased, management of CVC and HLB have shown good results in the last years. According to Fundecitrus,14 CVC is practically extinct in the “citrus park” of Sao Paulo, while the HLB incidence remains stable. In Sao Paulo State, the largest citrus producer in the world, quarantine measures based on exclusion and eradication protocols are applied to manage CCK since the first outbreaks of the disease in 1957.19 However, over time the rules governing eradication have been changed, sometimes tightened, other times relaxed. An excellent review of the exclusion and eradication program in Sao Paulo state applied between 1999 and 2009 was recently published.20
The current legislation in the State of São Paulo (Resolution SAA - 147 of 2013) is the mildest of all times. The resolution states that contaminated trees should be eliminated and those within a 30-m radius should be sprayed with cupric fungicide, and sprays should be repeated at each new budding. The citrus growers must carry out quarterly surveys and provide semi-annual reports to the Secretary of Agriculture, as it had already been established for HLB.14 Every year, considerable increases in disease incidence have been noticed, although the currently available data are mathematical projections, since surveys are no longer carried out, and the disease situation was declared as epidemic.14
Xanthomonas citri
The first description of CCK was made in 1915, being the causal agent classified as Pseudomonas citri.21 Subsequently, it was reclassified in the species Xanthomonas axonopodis.22 Several forms of the disease (A, B, C, D and E) have been described according to the host range and symptoms caused in different citrus species (Table 2).22-25 The firstly described Asian CCK, caused by the Asian strains, is the most widespread and causes the most severe disease symptoms.25
Form |
Geographic distribution |
Pathovar |
Disease/Susceptible host |
Reference |
A |
Asia, South America, Oceania and the USA |
citri |
Asian canker / Sweet orange, mandarin, sweet lime, grapefruit and pummelo |
23-25 |
B |
South America |
aurantifolii |
Cancrosis B / Mainly affects lemons, but is also observed in common lime, sour orange, Volkamer lemon, lime, sweet lime, cider and occasionally mandarin and grapefruit |
|
C |
Brazil |
aurantifolii |
Mexican lime cancrosis / Mainly affects common lime, but is also observed in sour orange and lemon |
|
D |
Mexico |
aurantifolii |
Citrus bacteriosis / Common lime |
|
E |
Florida |
citrumelo |
Citrus bacterial spot (or Florida nursery canker) / Predominantly Swingle citrumelo |
Table 2 Described forms of bacterial citrus diseases caused by Xanthomonas citri
Several authors have attempted to classify the Xanthomonas strains that cause the different forms of disease in citrus at the species and subspecies level.13,22-25 But, to date, most proposed names are not widely accepted. In light of current knowledge, it can be said that CCK is caused by bacterial strains belonging to two main phylogenetic groups: Asian and South American, although there exist differences in host range as well as phenotypic and genotypic variation within groups.13,14,16,19,21-24 Because of the need for an appropriate name for a quarantine pathogen, X. citri is used to refer to any Xanthomonas that causes CCK symptoms. The strains that cause the different forms of CCK are often referred to (and widely accepted) as pathovars citri (canker "A"), aurantifolii ("B", "C" and "D") and citrumelo ("E") (Table 2).13,22-25
Xylella fastidiosa
X. fastidiosa was first reported in 1884 as the causal agent of Pierce’s disease of grapevine in Southern California, and validly named in 1987.26 It has been estimated that Xfa can colonize more than 300 plant species belonging to 163 families.27 The ability of Xfa to infect a wide range of plant species has been attributed to a high rate of acquisition of genetic material through natural transformation and intra-specific recombination.28-30 The Xfa species is endemic to America, but its distribution has rapidly expanded to the Asian and European continents, most likely due to exchange of contaminated plant material.30-32
Aspects of the taxonomy, biology, and pathogenicity of Xfa remained obscure for some time due to the inability to grow it in culture medium. Sequencing of its whole genome33 and the possibility to obtain cultures in artificial medium34,35 were instrumental for the advancement of our knowledge about this bacterial plant pathogen. Studies on the genotypic and phenotypic diversity of Xfa have historically provided confusing and inconclusive results,30,36,37 likely due to: (i) the small number of isolates used in phylogenetic studies, which are normally obtained from a small number of host species and a few geographic locations; (ii) the diversity of methods used by different researchers, which makes it difficult to carry out reliable comparisons; and, (iii) the natural competence of Xfa, which directly influences gene flow, and consequently, the evolution of the species.28-30
Some of the problems related to the taxonomy of Xfa were solved due to the application of Multi-Locus Sequence Typing (MSLT).30,38,39 However, there exists controversy on the number of Xfa subspecies. According to Almeida and Nunney,30 four Xfa subspecies are currently accepted based on geographic origin and host specificity: fastidiosa, multiplex, sandyi, and pauca. In addition, the subspecies tashke and morus were proposed (Table 3).31,37-42 Finally, strains associated with pear leaf scorch in Taiwan have been shown to belong to a different Xfa genotype based on the sequence of the 16S rDNA region.30,43,44 Despite subspecies pauca causing disease in both coffee and citrus, there seems to be host specialization within the subspecies, since strains from coffee cannot infect citrus and vice versa.41
Subspecies |
Geographic Distribution |
Main Diseases and Hosts |
Reference |
fastidiosa |
North, Central, and South America |
Pierce's disease of grapevine and almond leaf scorch; also infects almond trees, alfalfa, and maple |
|
morus |
Eastern USA |
Mulberry leaf scorch |
37-42 |
multiplex |
Subtropical North America and South America |
Decline of different types of trees, including peach and plum tree |
|
pauca |
Central and South America, Europe |
Coffee leaf scorch and citrus variegated chlorosis |
|
sandyi |
Southern USA |
Oleander leaf scorch |
|
tashke |
Southwestern USA |
Isolated from Chitalpa tashkentensis |
Table 3 Proposed subspecies of Xylella fastidiosa
Recent studies using genome-wide comparisons confirmed the difficulty to conclusively classify Xfa strains. For instance, Marcelletti and Scortichini,45 used genome-wide average nucleotide identity (ANI) and tetranucleotide frequency correlation coefficient analyses, as well as phylogeny based on genome-wide sequences and sequences of 956 core gene families to compare 21 strains from different geographic origins and host plants and found three well-defined phylogenetic groups corresponding to the subspecies fastidiosa, multiplex and pauca. Strains previously proposed to be classified as subspecies sandyi and morus clustered together with strains of subspecies fastidiosa. In contrast, Giampetruzzi et al.32 conducted phylogenetic analysis with 27 Xfa strains using genome-wide SNPs and pangenome sequences and found results consistent with the separation of strains into five subspecies: fastidiosa, multiplex, pauca, sandyi, and morus.
Candidatus liberibacter
Although HLB, also known as Citrus greening, had been known in China for more than 100 years,6,46 the bacterial nature of its causal agent was only known in the 1970s.6,47 Several independent groups reported cultivation of CaL species in artificial media48-50 but their results have not been reproducible and consistent, and to date (despite considerable effort), there has been no success in culturing strains belonging to CaL species that cause disease in citrus. Jagoueix et al.8 based on 16S rDNA sequence comparisons, showed that the citrus pathogens belonged to the alpha sub-division of proteobacteria and allocated them in the provisional taxon Candidatus, in accordance with the accepted proposal to classify incompletely described prokaryotes.51 Accordingly, Jagoueix et al.8 proposed Asian strains to be classified as ‘Ca. Liberibacter asiaticus’ and African strains as ‘Ca. Liberibacter africanus’. Since then, the 16S rDNA sequence has become a molecular marker widely used for classification of CaL species. In 2004, a new species causing HLB was identified in Brazil and, based on 16S rDNA sequence comparisons, named ‘Ca. Liberibacter americanus’.10
In addition to citrus, CaLaf strains have been isolated from rutaceous trees in South Africa. Such strains show genetic divergence from citrus strains as assessed by phylogenetic analyses based on 16S rDNA, outer membrane protein (omp), and 50S ribosomal (rplJ) gene sequences. As a consequence, five CaLaf subspecies have been proposed: subspecies ‘capensis’,9 ‘clausenae’, ‘vepridis’, ‘zanthoxyli’,52 and ‘tecleae’.53 The implications of the association of CaLaf with rutaceous plants and their allocation in different subspecies for the biology of the pathogen, such as host specificity, vector transmission, disease dispersal or virulence remain to be investigated.
Genome sequencing of several strains of CaL revealed a strong genome-size reduction, most likely reflecting their intimate biotrophic association with the host plant.54-57 Recently, the bacterial isolate BT-1 was recovered in pure culture from the phloem sap of defoliating mountain papaya (Carica stipulata × C. pubescens) in Puerto Rico and shown to be closely related to CaLam and CaLas based on similarity of 16S rDNA and conserved protein sequences as well as on genome-wide ANI analysis.58,59 The isolation and genome sequencing of this isolate, classified as Liberibacter crescens (Lcr), allowed for the first time a comprehensive description of the morphological, biochemical, and physiological characteristics of the genus Liberibacter.59
Although no host plant for it has been identified and its genome has not suffered a strong size reduction as in the CaL spp.,54-59 Lcr has become an important model to investigate biological aspects that could be relevant to grow in vitro and to control CaL species. Fagen et al.57 used RAST,60 KEGG61 and manual annotation to compare the genome sequence of Lcr BT-1 with those of CaLas psy62 and ‘Ca. Liberibacter solanacearum’ ZC1, the causal agent of tomato psyllid yellows and potato zebra chip, and found a total of 207 genes of known functions and 238 hypothetical genes of unknown function unique to Lcr, some of which could contribute to its ability to grow in vitro. The CaL species lost the ability to synthesize proline, phenylalanine, tryptophan, cysteine, tyrosine and histidine as well as components of several systems that could compromise their ability to sense and adjust to environmental fluctuations, such as two-component regulatory systems, the stringent response, and an alternate cytochrome pathway.59 In addition, Lai et al.62 using genome-wide transposon mutagenesis identified 314 protein-coding genes required for Lcr BT-1 growth in BM7 medium. Seventy-six of the 314 essential genes have no homologs in CaL, which could be targeted for further research aimed to understand why the plant pathogens cannot grow in that particular medium. Nonetheless, the requirement of the identified genes for CaL species growth in culture media has yet to be elucidated. The remaining 238 genes have homologs in CaL, and the authors suggest that could be targets for development of control drugs against the plant pathogens.4,7-14,20,21,25
Secretion systems and effectors
Xcc, Xfa, and CaL spp. colonize different citrus tissues. Xcc survives epiphytically, penetrates the host leaves through natural openings and wounds, and colonizes the mesophyll cells. Xfa and CaL spp., are transmitted by insect vectors and colonize the xylem and phloem vessels, respectively.4,7-14,20,21,25 Among them, only CaL spp. colonizes the intracellular space, strictly infecting the phloem sieve tubes.4-8,12,14 Not surprisingly, the molecular mechanisms that they utilize to cause disease in citrus plants are different. Genes coding for transporters, structural components of secretion systems, transcriptional regulators, and synthesis of structural molecules that provide protection to the bacterial cell have been suggested to be pathogenicity or virulence factors. In this review, we will mostly discuss about proteins that are secreted to the host tissue in order to promote bacterial parasitism and pathogenesis.
The Type I secretion system (T1SS) is present in all three citrus bacterial pathogens. It is important for bacterial protection and competition as well as for pathogenesis. During pathogenesis, the T1SS secretes diverse types of proteins, including proteases, hemolysins, bacteriocins and effector proteins.63,64 In CaL spp., like in other Gram-negative phytopathogenic bacteria, there are multiple copies of the gene encoding the TolC protein, an outer membrane protein required by both efflux systems and the T1SS. In contrast, Xfa strains that cause CVC have only one copy of tolC, despite having multiple T1SS.65 TolC allows passage of substrates from the bacterial cytosol to the external milieu.64 The contribution of effectors secreted by the T1SS to virulence of strains causing CCK, CVC, and HLB remains to be investigated.
X. citri has two type II secretion systems (T2SS), coded by two different gene clusters: Xps and Xcs.66 The T2SS is involved in transport of sugars, secretion of enzymes for cell-wall degradation, including hemicellulases, glucanases, pectinases and polygalacturonases, as well as in absorption of nutrients released from such a degradation.67 The contribution of enzymes secreted by the T2SS to the virulence of Xcc has been demonstrated. For instance, expression of the pectate lyase gene pel1 from Xcc strain XW19 (which causes lesions with water-soaked margins) in strain XW121 (which does not cause water-soaked margins) conferred to the transformants the ability to cause water-soaked margins when inoculated onto grapefruit plants.68 Homologous of pectolytic genes have been identified in diverse Xcc strains. It has also been shown that a pel3 deletion mutant of strain XW19 displays reduced growth and induces smaller canker lesions on leaves of Mexican lime.69 Also, deletion of the bgl3 gene, coding for an endoglucanase, in strain Xcc 20-1 caused delay of CCK symptoms development.70
It is believed that the Xfa T2SS is mainly associated with degradation of the pit membranes, facilitating bacterial colonization, mobilization of nutrients and subversion of plant defense responses.71,72 No genes coding for extracellular cell-wall degrading enzymes were found in the genome of CaLas. However, genes coding for proteins of the T2SS general secretion pathway, as well as for proteins putatively involved in the formation of type IV pili (T4P) are present.4,54 T4P are flexible surface filaments that are assembled through the general secretion pathway and play diverse roles, for example, in twitching motility, surface adhesion, natural transformation, biofilm formation and chemotaxis.73,74 It has been speculated that they may provide an important function in auto-aggregation and biofilm formation for CaLas,54 but this hypothesis remains to be proved. In Xfa, T4P are required for obstruction of sap flow, intercellular communication, and migration of bacterial cells within the xylem.75-78 In Xcc, it was recently shown that, like in Xfa, T4P are required for twitching motility, adhesion, and biofilm formation.74,79
The Sec-dependent secretome was predicted from the genome sequences of several CaL species and Lcr. Eighty-six out of 166 predicted proteins were validated to contain signal peptides when expressed in fusions with alkaline phosphatase (phoA) in Escherichia coli.80 Also, 16 effectors were predicted from the genome sequence of CaL Psy62 as proteins that had a signal peptide, lacked transmembrane domains, had a molecular size ≤ 250 amino acids, and showed no sequence similarity to proteins of known function. Predicted effector Las5315, which targeted the chloroplast, caused cell death in N. benthamiana and was associated with callose deposition and hydrogen peroxide accumulation.81
The main factors used by Xcc to cause disease in citrus are proteins of the PthA effector family, whose coding genes are often present in multiple copies in the genome.82 PthA belongs to the so-called AvrBs3/PthA protein family. Members of the AvrBs3/PthA family are TAL (transcription activator-like) effectors, which are translocated by the type three secretion system (T3SS) from the bacterial cytoplasm directly to the interior of host cells where they bind to sequences called effector binding elements (EBE) in the promoter of host susceptibility genes to activate their expression.83 The main susceptibility target of PthA in C. sinensis is LOB1 (Lateral organ boundaries 1), a member of the LBD (LOB Domain) family of proteins, which are regulators of plant organ development.84 Homologs of phtA have been found in strains belonging to Xcc groups that show differences in host range and geographic distribution,85 all of them were shown to induce the expression of CsLOB1 in grapefruit and sweet orange.86 The number and type of LOB1 alleles differ among citrus genomes. For instance, Duncan grapefruit has two allele types in a ratio 1:187 and Wanjincheng orange has two in a ratio 3:1,88 whereas Satsuma mandarin and Chandler pummelo harbor only one.89,90
Some effectors secreted by the T3SS are recognized by plant resistance proteins eliciting a response known as ETI (effector triggered immunity), considered to be the second layer of defense after PTI (PAMP-triggered immunity) in the plant immune system. Effector proteins recognized by the plant surveillance system confer an avirulent phenotype to the pathogen, and hence, they are called avirulence (Avr) proteins. The recognition of the effector culminates in a programed cell death known as the hypersensitive response (HR).91 Avirulence proteins are main determinants of the pathogen host range. A genome comparison revealed that the effector genes xopAF and avrGf1 are present in Xcc Xcaw12879 (restricted to Mexican lime), but are absent in Xcc XccA306 (type A; broad host range). Analyses of xopA1 and avrGf1 single and double mutants revealed that AvrGf1 causes the HR in Duncan grapefruit and XpAF contributes to virulence in Mexican lime.92 These results indicate that AvrGf1 restricts the host range of Xcc toward Duncan grapefruit.
Xcc and Xfa have type IV secretion system (T4SS) gene clusters similar to that of the Bordetella pertussis involved in the secretion of pertussis toxin.93 The T4SS is associated with the transport of macromolecules from the bacterial cytoplasm to the host cytoplasm. In CaLas the T4SS is absent.4,54 The participation of the T4SS in the pathogenicity of Xcc and Xfa is yet to be determined.
During the pathogenic process, bacterial cells attach onto host surfaces and develop communities with the capacity to cause infection. Biofilm formation is important for the pathogenicity of Xcc and Xfa.94,95 The genomes of Xfa and Xcc contain genes coding for adhesins predicted to be secreted by mechanisms resembling type V secretion systems (T5SS). Adhesins are high molecular weight proteins that play key roles in adhesion and biofilm formation.54,96 Genes coding for adhesins are up-regulated in Xfa biofilms in vitro.94 In Xcc, mutation of the gene encoding the filamentous hemagglutinin-like protein XacFhaB, which presumably is part of a two-partner T5SS, impairs leaf surface attachment and biofilm formation. The XacFhaB mutant caused more dispersed and fewer canker lesions compared to the wild type strain.97 Xfa has some high molecular weight proteins that show sequence similarity to XacFhaB, although restricted to the N- and C-terminal portions of the protein.93 The T5SS is also responsible for secreting other class of proteins that are involved in virulence. In CaLas, the T5SS-secreted proteins LasAI and LasAII were recently shown to be virulence factors targeting the host cell mitochondrion. LasAI and LasAII have structural characteristics of leucine-rich repeat proteins and mitochondria-targeting autotransporters.97
Perspectives for disease control
Physical, chemical and biological approaches: Currently, there is no citrus varieties resistant to HLB, no cure for the disease has been identified, and it has not been eradicated from any area.20 Although thermotherapy has been used to eliminate pathogens from infected plant material for decades, and it was long ago shown that exposure of budwood and seedlings to different regimes at 50 ̊C eliminates HLB symptoms,98,99 there is a resurgent interest in investigating its potential use for controlling the disease. Recent studies have demonstrated that exposure of unhealthy Citrus reticulata, C. paradisi, and C. limon plants to temperatures between 40 ̊C and 48 ̊C was highly efficient in reducing the titer of CaLas and HLB symptoms.100,101
The effectiveness of several bactericides against CaLas has been demonstrated in greenhouse studies.102,103 However, foliar application of bactericides suffers from several limitations, including rapid degradation of the compound under sunlight and risk of environmental contamination. CaL spp. are restricted to the phloem, which makes it difficult to effectively control them using foliar applications of bactericides. Among the alternatives that have been shown to overcome these limitations are trunk injection and the use of nanoemulsion formulations that enhance bactericide penetration.104,105
Interestingly, Pagliai et al.106 showed that targeted inhibition of the transcriptional activator LdtR, involved in the regulation of a transpeptidase encoded by the gene ldtP, could be an approach for developing therapeutic molecules against CaLas. The authors screened two libraries containing more than 1300 small molecules and found that phloretin, hexestrol, and benzbromarone inhibited binding of LdtR to DNA. In in vitro assays, the small molecules caused morphological changes and sensitivity to osmotic stress in CaLas cells similar to those caused by mutations of ldtR in Sinorhizobium meliloti and Lcr, indicating that these compounds have potential use in the control of HLB.
Copper-based bactericides are routinely applied in several growing areas to control CCK. However, the extensive use of copper compounds can lead to soil contamination and selection of copper-resistant Xcc strains.107 On the other hand, antibiotics have not been as effective as copper compounds for controlling CCK because of their limited residual activity on foliar surfaces.108 Hence, there is an urgent need to find alternative control strategies. Significant reductions in the percentage of infected leaves and disease intensity were obtained by application of phages isolated from infected leaves or their combination with acibenzolar-S-methyl (ASM) onto Mexican lime (C. aurantifolia) plants previously inoculated with Xcc under greenhouse and field conditions.109 Similar results had previously been reported.110
Application of Pseudomonas protegens CS1, isolated from the surface of lemon leaves (C. limon cv. Limoneira 8A) provided protection against CCK. The active compound responsible for bacterial inhibition was identified as enantio-pyochelin, which when applied to the leaves also caused attenuation of disease symptoms. It was shown that one of the mechanisms by which P. protegens CS1 protects the citrus plant is by an increase in Xcc lipid peroxidation due to the generation of reactive oxygen species by enantio-pyochelin.111 Also, reductions in bacterial populations and/or disease symptoms were observed after application of the biofilm-inhibiting compounds D-leucine, 3-indolylacetonitrile112 and bismerthiazol to inoculated Duncan grapefruit plants. Bismerthiazol, which has been used to control rice bacterial blight, not only inhibits bacterial growth but also induces plant resistance responses.113
With regard to Xfa, an interest in using endophytes to control it has emerged because of the observation that some endophytes are frequently associated with infected citrus trees and occupy the same xylem locations as the pathogen.14,115 Among such endophytes, Methylobacterium mesophilicum and Curtobacterium flaccumfaciens have been shown to cause reduction in Xfa populations and disease symptoms in co-inoculation experiments using the model plant Catharanthus roseus .116,117 A review on the possible alternatives to use these endophytes in the control of CVC was recently published.118
Plant genetic resistance
Regarding the search for alternatives using plant resistance, Ramadugu et al.119 found resistant accessions of Australian desert lime (Eremocitrus glauca), curry leaf (Bergera koenigii), Japanese prickly ash (Zanthoxylum ailanthoides), Hesperethusa (Naringi crenulata), orange jasmine (Murraya paniculata), and white sapote (Casimiroa edulis), in which the pathogen replicated transiently, but could not establish in the plant. The authors suggested that accessions of Eremocitrus and Microcitrus (ranked as tolerant), which are sexually compatible with Citrus, have the potential for the generation of HLB resistant cultivars. The results of that study support previous findings indicating the existence of resistance in species closely related to Citrus.120,121
A recent report showed that application of several plant defense inducers, such as ß-aminobutyric acid (BABA), 2,1,3-benzothiadiazole (BTH), and 2,6-dichloroisonicotinic acid (INA), singly or in combination suppressed CaLa growth in planta and the progress of HLB symptoms.122 Activation of the brassinosteroid and jasmonic acid pathways also seem to provide some level of protection against HLB. Canales et al.123 demonstrated that spraying 24-epibrassinolide (eBL) to Mexican lime (C. aurantifolia) plants under greenhouse conditions and to Valencia sweet orange (C. sinensis) plants in the field caused significant reductions in CaLas titers compared to non-sprayed control plants. Application of eBL caused the induction of genes involved in several defense-response pathways.
Transgenic and gene overexpression approaches have been used in an attempt to control bacterial citrus diseases. Examples of transgenes or genes whose overexpression confer enhanced resistance are shown in Table 4.124-138 Nonetheless, a main limitation of the transgenic approach is that adoption of genetically modified plants takes many years because of the rigorous field testing required and the long regulation process.
Gene |
Function |
Source |
Type of Expression |
Pathogen* |
Citrus Plant |
Reference |
attA |
Attacin A; antimicrobial peptide |
Tricloplusia ni |
Transgenic |
CaLas, Xcc |
Natal, Pera, and Valencia sweet orange |
124-126 |
CB |
Cecropin B; lytic peptide with antibacterial activity |
Chinese tasar moth (Antheraea pernyi) |
Transgenic |
CaLas |
Tarocco blood orange (Citrus sinensis) |
|
FLS2 |
Plant recognition receptor that recognizes bacterial flagellin |
Nicotiana benthamiana |
Transgenic |
Xcc |
Hamlin sweet orange and Carrizo citrange |
|
FLS2 |
Plant recognition receptor that recognizes bacterial flagellin |
Fortunella margarita and C. reticulata |
Transient expression |
Xcc |
Duncan grapefruit |
|
MAF1 |
RNA polymerase III repressor |
Sweet orange (C. sinensis) |
Overexpression |
Xcc |
Hamlin sweet orange |
|
NH1 |
A, thaliana NPR1 ortholog |
Pummelo (C. maxima) |
Overexpression |
Xcc |
Duncan grapefruit |
|
NPR1 |
Non-expressor of PR genes 1; master regulator of the SA pathway |
Arabidopsis thaliana |
Transgenic |
CaLas, Xcc |
Hamlin and Valencia sweet orange, and Duncan grapefruit |
132-134 |
rpfF |
Synthesis of a diffusible signal factor (DSF) involved in quorum sensing |
Xylella fastidiosa |
Transgenic |
Xcc |
Hamlin, Natal, Pera, and Valencia sweet orange |
|
STS3-1 |
Linalool synthase |
‘Miyagawawase’ satsuma mandarin (C. unshiu) |
Overexpression |
Xcc |
Hamlin sweet orange |
|
thi |
Thionin; cysteine-rich antimicrobial peptide |
Synthetic |
Overexpression |
CaLas, Xcc |
Carrizo citrange (C. sinensis x Poncirus trifoliata) |
|
Xa21 |
Plant recognition receptor |
Rice (Oryza longistaminata) |
Transgenic |
Xcc |
Hamlin, Natal, Pera, and Valencia sweet orange |
Table 4 Examples of genes whose expression in citrus plants confer resistance against citrus canker and/or Huanglongbing
*CaLas, ‘Candidatus Liberibacter asiaticus’; Xcc, Xanthomonas citri
Gene edition has also been used to procure resistance against CCK. In an attempt to block the activation of CsLOB1, the EBE of the type I allele recognized by PthA4 was edited using Cas9/sgRNA technology in Ducan grapefruit, but resistance to CCK was not observed.87 Since grapefruit resulted from hybridization between sweet orange and pummelo (C. maxima),139 it contains CsLOB1 alleles from both sweet orange (type I) and pummelo (type II). Later, Peng et al.88 showed that Cas9/sgRNA-mediated editing of the EBEs of both CsLOB1 alleles in Wanjincheng orange (C. sinensis) conferred high levels of resistance against CCK. Reduced susceptibility was also obtained when the coding sequence of CsLOB1 in Duncan grapefruit was edited using Cas9/sgRNA technology.140 No apparent effect on plant morphology and development was observed in the edited plants, indicating that genome editing of susceptibility genes is a promising alternative to obtain plants resistant to CCK.
The taxonomy of Xcc, Xfa, and CaL spp. is still controversial and it is expected that new proposals for allocation of strains in new species or subspecies appear in the near future. In Brazil, Xcc is the species that currently causes the most severe damage. After several decades, it re-emerged as the main bacterial disease of citrus as a result of inefficient management practices. The interaction of citrus species with these bacterial pathogens is one the best examples to understand the different strategies that bacteria utilize to infect their host plants. Whereas for Xcc the T3SS plays a major role in pathogenicity, Xfa and CaL lack such a secretion system and seem to depend on genes involved in adhesion and biofilm formation. The management strategies applied for CCK, CVC, and HLB are very similar, except for an efficient control of insect vectors required in the cases of CVC and HLB. New alternatives to control the pathogens are emerging as a result of research that demonstrate the antibacterial activity of new compounds and the enhanced resistance of transgenic plants. Among the proteins with potential use in obtaining transgenic plants resistant to bacterial citrus diseases are plant recognition receptors, master regulators of the SA pathway, cercopins, and thionins. Notably, CRSPR/Cas9 genome editing of TAL susceptibility genes (either EBE or coding sequence) was proved to confer enhanced resistance against Xcc.
We thank financial support provided by the Brazilian agencies CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior), CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) and FAPEMIG (Fundação de Amparo à Pesquisa de Minas Gerais) to conduct research in our laboratories.
The authors declare that they have no conflict of interest and that financial support provided by funding agencies did not influence the content of this article.

©2017 Mendonça, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.
 World Immunization week is celebrated in the last week of April. This highlights the importance of vaccination to all age groups to protect from diseases. So, we are enthusiastic to invite all to submit articles which help our readers in raising the awareness on this vaccination for vaccine-preventable diseases. You can also avail a best discount of 35% for your submissions in our Journal of Bacteriology & Mycology: Open access (JBMOA).
World Immunization week is celebrated in the last week of April. This highlights the importance of vaccination to all age groups to protect from diseases. So, we are enthusiastic to invite all to submit articles which help our readers in raising the awareness on this vaccination for vaccine-preventable diseases. You can also avail a best discount of 35% for your submissions in our Journal of Bacteriology & Mycology: Open access (JBMOA).