eISSN: 2378-315X


Research Article Volume 14 Issue 2
Biostatistics and Data Science Program, LSU Health Sciences Center, USA
Correspondence: Evrim Oral, Biostatistics and Data Science Program, LSU Health Sciences Center, New Orleans LA, USA
Received: September 15, 2025 | Published: October 1, 2025
Citation: Mohamed M, Vaidya L, Mamnan M, et al. Time-series modeling of COVID-19 cases in the United States with google search trends. Biom Biostat Int J. 2025;14(2):61-66. DOI: 10.15406/bbij.2025.14.00435
Syndromic surveillance offers a rapid, low-cost approach to monitoring emerging health threats, complementing traditional case-based systems. This study investigates the utility of Google Trends data as a proxy for incident COVID-19 cases in the United States between March 2020 and April 2023. Weekly search interest for terms including covid, COVID-19, fever, mask, flu, and COVID-19 vaccine was analyzed alongside reported cases. Time-series modeling compared vector autoregressive (VAR), transfer function (TFM), and web-search-only (WSO) approaches. VAR models produced the most accurate forecasts of weekly cases and epidemic peaks, while TFM showed moderate accuracy, and WSO models—although overestimating magnitudes— were useful in identifying epidemic onset, peak timing, and decline. Our findings highlight the promise of integrating web-based search data into surveillance frameworks, especially in settings with limited diagnostic or reporting capacity, while also underscoring limitations such as news bias, confounding from overlapping symptoms, and the need for early calibration in novel outbreaks.
Traditional surveillance systems form the foundation of public health monitoring and rely primarily on case-based reporting and laboratory confirmation to track disease incidence. Physicians, hospitals, and laboratories are required to report notifiable diseases to local, state, and national health authorities, ensuring high specificity and standardized case definitions across regions. These systems provide accurate and detailed information on confirmed cases, including demographic and clinical characteristics, which supports in-depth epidemiologic analyses.
However, traditional surveillance often suffers from delays due to the time required for diagnostic testing, reporting, and data compilation, making it less effective for early outbreak detection. Underreporting of mild or untested cases and the need for significant laboratory and personnel resources further limit its timeliness and coverage. In addition to these systemic challenges, barriers such as delays in developing and disseminating accurate diagnostic tests, social and psychological factors (e.g., stigma, misinformation, or hesitancy to seek testing), and logistical constraints (e.g., limited availability or accessibility of testing sites) can also impede timely detection. While essential for producing official case counts and monitoring long-term trends, traditional surveillance systems are constrained in their responsiveness, underscoring the need for complementary approaches such as Syndromic surveillance to enhance early detection and support timely epidemic control. Syndromic surveillance addresses many of these limitations by collecting and analyzing health-related data prior to laboratory confirmation, thereby providing earlier signals of emerging health threats.1 Rather than depending on confirmed diagnoses, Syndromic systems monitor indicators such as emergency department visits, over-the-counter medication sales, school absenteeism, call center data, and digital sources including internet search trends and social media activity. By identifying unusual patterns in these data streams, Syndromic surveillance can detect potential outbreaks days or weeks earlier than traditional systems, allowing for timelier public health response. Although less specific than laboratory- confirmed surveillance, its ability to enhance situational awareness and provide early warnings makes Syndromic surveillance a valuable complement to traditional systems, particularly during rapidly evolving epidemics such as COVID-19.
Since its introduction in the 1990s, the application of Syndromic surveillance has expanded considerably and, in many instances, has enabled accurate predictions of disease trends and outbreak dynamics.2 In contrast to traditional surveillance systems, which depend on laboratory-confirmed diagnoses, syndromic surveillance offers a less resource- intensive approach and has demonstrated the ability to identify outbreaks earlier.3,4 It has also been applied to predict the progression of epidemics, including the timing and magnitude of peak case incidence5,6 and is particularly valuable in settings where laboratory testing capacity is constrained.7 Collectively, these advantages enhance the timeliness of outbreak detection and response, thereby reducing the risk of rapid increases in case incidence and mitigating pressure on healthcare systems. Edge et al.,8 among the earliest studies in this area, analyzed over-the-counter medication sales from a single retailer and successfully detected gastrointestinal infections in near real-time, outperforming traditional surveillance systems, which confirmed only 10% of epidemic cases. Notably, in the context of COVID-19, Syndromic surveillance systems have demonstrated the capacity to predict outbreaks 1–2 weeks in advance with high accuracy.3,9,10 Mahmud et al.,3 using self-reported symptoms, demonstrated the ability to predict COVID-19 trends in Bangladesh 1–2 weeks earlier than traditional approaches. Similarly, Vigfusson et al.4 investigated H1N1 influenza using call record data and identified the potential for six-day earlier predictions.
Digital data sources, such as Google Trends, have further advanced syndromic surveillance research. Several studies have shown that peak search interest for COVID-19 occurred 11–17 days prior to peaks in confirmed cases.11- 13 Effenberger et al.11 reported that epidemic peaks across multiple countries could be predicted up to 11.5 days in advance based on search activity.
Lampos et al.12 examined the predictive value of different COVID-19–related search terms, noting that some terms (e.g., covid) showed stronger correlations with case counts than others (e.g., covid-19), while certain terms (e.g., flu-like symptoms) demonstrated inverse correlations, suggesting the influence of confounding factors.
Methodologically, most studies have relied on moving average models to account for weekly seasonality in COVID-19 testing patterns and to align with epidemic threshold definitions.3,8,13 Correlation analysis has been widely used to evaluate associations between search terms and case counts, as well as to identify optimal lag structures for prediction.11, 12
The present study builds upon this body of research by demonstrating that Google search trends can be used to accurately estimate the number of COVID-19 cases in the United States, identify peak case counts, determine optimal lag periods for prediction, and quantify prediction errors. Through time series analysis, we evaluate the relationship between Google search trends and COVID-19 case incidence, with particular attention to how this relationship may have been influenced by the introduction of vaccination programs.
Data were collected from Google Trends using the following search terms: COVID-19, covid, fever, mask, flu, and COVID-19 vaccine. Google Trends data are scaled from 0 to 100, where 100 indicates the highest search popularity for a given term, and 50 represents half that level. Daily COVID-19 case counts were obtained from the World Health Organization COVID-19 tracking program. Vaccination data, including counts and population percentages for first doses, full vaccination, and booster doses, were obtained from the Centers for Disease Control and Prevention website. The study period spanned March 2020 through April 2023.
New cases and vaccination data were aggregated into weekly values to align with the Google Trends data, resulting in a dataset spanning 166 weeks. For visual comparison, new cases were scaled to their maximum values; however, the scaled data were not used for model building or prediction.
All statistical analyses were conducted in R (version 4.3.2). Descriptive statistics were first calculated for the Google search terms listed above, along with weekly new cases, first vaccine doses, completed vaccinations, booster counts, and corresponding population percentages. Correlation analyses were performed for the full dataset, and correlation matrices were compared before and after reaching 70% first-dose vaccination coverage. The data were then divided into training (80%) and testing (20%) sets. We applied three approaches—the Vector Autoregressive (VAR) model, Transfer Function Model (TFM), and Web-Search-Only (WSO) model—to assess new COVID-19 cases. Stationary was evaluated using the Augmented Dickey–Fuller test, and model performance was assessed using mean squared error (MSE).
Descriptive statistics are presented in Table 1. Correlation analyses between Google search terms and new cases showed the strongest statistically significant association with the term covid (Figure 1). This association persisted after stratifying the population by vaccination status (<70% vs. ≥70% first-dose coverage), with a notably higher correlation coefficient at high vaccination levels (r = 0.957, p < 0.001) (Figure 1&2).
|
𝝆 (p-value) |
New Cases |
COVID- 19 Vaccine |
Flu |
Mask |
Fever |
covid |
COVID- 19 |
|
COVID-19 |
0.144 |
0.304 |
0.501 |
0.749 |
0.439 |
0.553 |
1 |
|
-0.064 |
(<.001) |
(<.001) |
(<.001) |
(<.001) |
(<.001) |
(<.001) |
|
|
covid |
0.657 |
0.78 |
-0.039 |
0.215 |
0.131 |
1 |
|
|
(<.001) |
(<.001) |
-0.616 |
-0.005 |
-0.094 |
(<.001) |
||
|
Fever |
0.337 |
-0.2 |
0.74 |
0.195 |
1 |
||
|
(<.001) |
-0.01 |
(<.001) |
-0.012 |
(<.001) |
|||
|
Mask |
-0.022 |
-0.04 |
0.36 |
1 |
|||
|
-0.781 |
-0.607 |
(<.001) |
(<.001) |
||||
|
Flu |
-0.036 |
-0.249 |
1 |
||||
|
-0.649 |
-0.001 |
(<.001) |
|||||
|
COVID-19 Vaccine |
0.212 |
1 |
|||||
|
-0.006 |
(<.001) |
Pearson correlation coefficients between google search trends for the terms COVID-19 vaccine, flu, mask, fever, and covid with new cases in the following week.
|
Vaccination below 70% |
||||||||
|
𝝆 (p-value) |
New Cases |
COVID- 19 Vaccine |
Flu |
Mask |
Fever |
covid |
COVID- 19 |
|
|
COVID-19 |
-0.058 |
-0.034 |
0.559 |
0.610 |
0.759 |
0.224 |
1 |
|
|
-0.588 |
-0.75 |
(<.001) |
(<.001) |
(<.001) |
-0.034 |
(<.001) |
|
|
|
covid |
0.586 |
0.84 |
-0.309 |
-0.246 |
-0.073 |
1 |
|
|
|
(<.001) |
(<.001) |
-0.003 |
(0.020) |
-0.495 |
(<.001) |
|
||
|
Fever |
-0.127 |
-0.268 |
0.846 |
0.379 |
1 |
|
||
|
-0.232 |
-0.011 |
(<.001) |
(<.001) |
(<.001) |
|
|||
|
Mask |
-0.301 |
-0.435 |
0.341 |
1.000 (<.001) |
|
|||
|
-0.004 |
(<.001) |
-0.001 |
|
|
||||
|
Flu |
-0.228 |
-0.433 |
1 |
|
||||
|
-0.031 |
(<.001) |
(<.001) |
|
|||||
|
COVID-19 |
0.218 |
1 |
|
|||||
|
Vaccine |
-0.039 |
(<.001) |
|
|||||
|
Vaccination above 70% |
||||||||
|
𝝆 (p-value) |
New Cases |
COVID- 19 Vaccine |
Flu |
Mask |
Fever |
covid |
COVID- 19 |
|
|
COVID-19 |
0.952 |
0.847 |
0.168 |
0.743 |
0.617 |
0.988 |
1 |
|
|
(<.001) |
(<.001) |
-0.147 |
(<.001) |
(<.001) |
(<.001) |
(<.001) |
|
|
|
covid |
0.957 |
0.802 |
0.218 |
0.675 |
0.684 |
1 |
|
|
|
(<.001) |
(<.001) |
-0.059 |
(<.001) |
(<.001) |
(<.001) |
|
||
|
Fever |
0.721 |
0.385 |
0.678 |
0.306 |
1 |
|
||
|
(<.001) |
(<.001) |
(<.001) |
-0.007 |
(<.001) |
|
|||
|
Mask |
0.686 |
0.597 |
0.114 |
1 |
|
|||
|
(<.001) |
(<.001) |
-0.325 |
(<.001) |
|
||||
|
Flu |
0.192 |
0.132 |
1 |
|
||||
|
-0.099 |
-0.257 |
(<.001) |
|
|||||
|
COVID-19 |
0.723 |
1 |
|
|||||
|
Vaccine |
(<.001) |
(<.001) |
|
|||||
Pearson correlation coefficients between google search trends for the terms COVID-19 vaccine, flu, mask, fever, and covid and new cases, before and after reaching 70% vaccination coverage, with correlations assessed for the following week.
|
Variable |
N |
Mean (SD) |
Minimum |
Maximum |
|
Google Search Terms |
||||
|
COVID-19 |
166 |
19.28 (17.06) |
2.22 |
100 |
|
covid |
166 |
28.08 (20.3) |
3 |
100 |
|
Fever |
166 |
36.717 (9.934) |
27 |
100 |
|
Mask |
166 |
21.83 (15.27) |
9 |
100 |
|
Flu |
166 |
13.95 (12.99) |
4 |
100 |
|
COVID-19 Vaccine |
166 |
17.03 (22.55) |
0 |
100 |
|
New Cases |
166 |
622086 (781481) |
0 |
5605477 |
|
Dose1 |
166 |
161768453 |
0 |
270214753 |
|
-113134233 |
||||
|
Dose1 Percent |
166 |
0.4878 (0.3412) |
0 |
0.8149 |
|
Completed Vaccination |
166 |
137285330 |
0 |
230632064 |
|
-99020716 |
||||
|
Completed Vaccination Percent |
166 |
41.4 (29.86) |
0 |
0.6955 |
|
Booster Doses |
166 |
49481830 |
0 |
118305291 |
|
-52985219 |
||||
|
Booster Percent |
166 |
0.1492 (0.160) |
0 |
0.3568 |
Table 1 Descriptive statistics for each variable
The autocorrelation and partial autocorrelation functions showed the strongest correlations at lags 1 and 2 (Figures 3a and 3b), consistent with an autoregressive model of order 2 (AR (2)). Cross-correlation analysis further indicated that the search term covid led new case counts by one week (Figure 3c) (Figure 3).
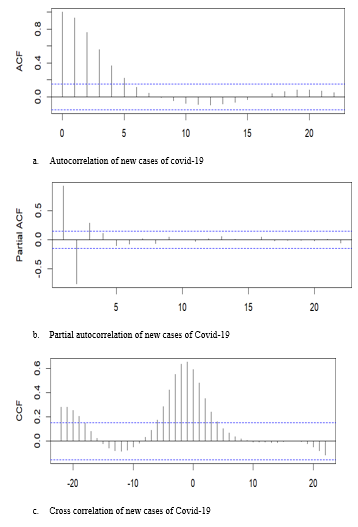
Figure 3 Autocorrelation (a), partial autocorrelation (b) and cross correlation (c) between google trend search term Covid and new Covid-19 cases.
For visual comparison, new cases were scaled to the 0–100 range used in Google Trends, with 100 representing the maximum and 0 the minimum. These scaled case values were then compared with search trends for the search terms COVID-19 and covid (Figure 4).
Stationary was assessed using the Augmented Dickey–Fuller (ADF) test, which indicated that new cases were stationary (DF = –3.8082, p = 0.02). For model development, 80% of the data (N = 133) were used for training and 20% (N = 33) for testing.
Vector auto regressive model (VAR)
The first model fit was a VAR model between the search term covid and new cases. The optimal VAR order was three lags, selected based on the Bayesian Information Criterion (BIC). The model coefficients are presented in Table 2, in the form.
|
Model |
Variable |
Estimate |
SE |
t value |
Pr(>|t|) |
|
New Cases |
covidt-1 |
1.23E+04 |
1.78E+03 |
6.898 |
2.46E-10 |
|
New Casest-1 |
1.86E+00 |
8.02E-02 |
23.12 |
< 2E-16 |
|
|
covidt-2 |
-8.42E+03 |
2.64E+03 |
-3.194 |
0.00178 |
|
|
New Casest-2 |
-1.32E+00 |
1.32E-01 |
-9.985 |
< 2E-16 |
|
|
covidt-3 |
-2.92E+03 |
2.03E+03 |
-1.437 |
0.15315 |
|
|
New Casest-3 |
4.05E-01 |
6.84E-02 |
5.927 |
2.89E-08 |
|
|
Const |
1.14E+04 |
2.70E+04 |
0.422 |
0.67395 |
|
|
Covid |
covidt-1 |
1.05E+00 |
9.01E-02 |
11.654 |
<2E-16 |
|
New Casest-1 |
6.24E-06 |
4.06E-06 |
1.536 |
0.1271 |
|
|
covidt-2 |
-5.13E-02 |
1.33E-01 |
-0.384 |
0.7016 |
|
|
New Casest-2 |
-1.35E-05 |
6.68E-06 |
-2.013 |
0.0463 |
|
|
Covidt-3 |
-6.12E-02 |
1.03E-01 |
-0.595 |
0.5531 |
|
|
New Casest-3 |
5.85E-06 |
3.46E-06 |
1.689 |
0.0938 |
|
|
const |
2.93E+00 |
1.37E+00 |
2.14 |
0.0344 |
Table 2 Vector auto regression model (VAR) coefficients
Where c is a constant vector, A1, A2, A3 are 2×2 coefficient matrices, and et is the error vector. Model performance was evaluated using 1-step-ahead and 2-step-ahead rolling forecasts on the 20% test dataset (Figure 5). The MSE was smaller for the 1-step forecast compared to the 2-step forecast (6.573 × 10⁹ vs. 13.075 × 10⁹), indicating reduced predictive accuracy over a two-week horizon (Table 2).
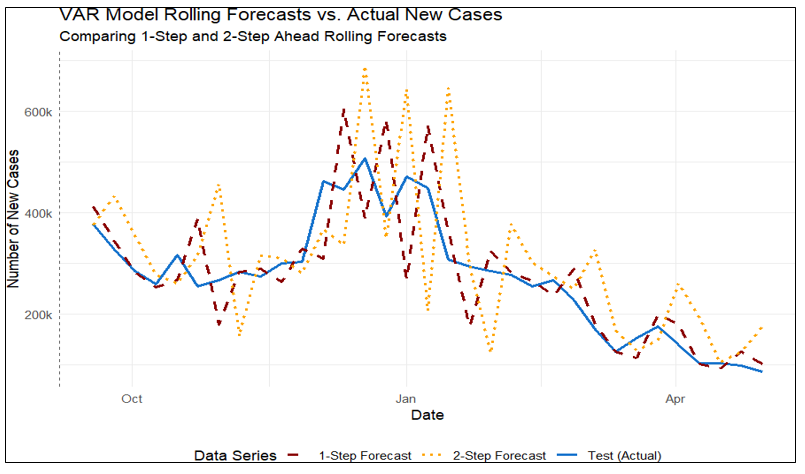
Figure 5 VAR model: time series plot of new cases with 1-step-ahead and 2-step-ahead rolling forecasts.
Transfer function model (TFM)
We also fit a TFM. Based on the autocorrelation functions (ACFs), partial autocorrelation functions (PACFs), and cross-correlation function (CCF) of covid and new cases (Figures 1–3), the selected model specified an ARIMA (2, 1, and 0) structure for new cases with the Google search term covid included as a covariate. The TFM coefficients are presented in Table 3, with the model.
|
Variable |
Estimate |
Std. Error |
|
Covidt |
1.23E+04 |
1.78E+03 |
|
New Casest-1 |
1.2161 |
0.0683 |
|
New Casest-2 |
-0.614 |
0.0674 |
Table 3 Transfer function model (TFM) coefficients
Where Δ describes the weekly difference, 𝛽 is the coefficient for Google search term covid, 𝜙1, 𝜙2 are the autoregressive coefficients, and e is the error term. Similar to VAR model, 1-step-ahead and 2-step-ahead rolling forecasts were performed, with the MSE smaller for the 1-step horizon (9.941 × 10⁹ vs. 25.913 × 10⁹; Table 3). However, the MSE for the TFM was larger than that of the VAR model, indicating less accurate predictions of weekly case counts and epidemic peak values. See Figure 6 for model performance on the test dataset (Table 3, Figure 6).
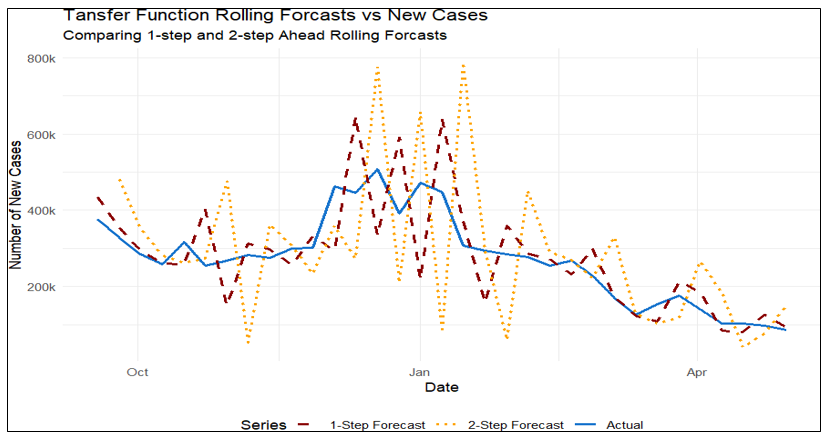
Figure 6 Transfer function model: time series plot of new cases with 1-step-ahead and 2-step- ahead rolling forecasts.
Web-search-only model (WSO)
The final model fit was the Web-Search-Only (WSO) model, which used Google search terms from the previous week to predict current COVID-19 case counts, without incorporating prior case values. This model was examined to assess its potential utility in situations where case data may be inaccurate or unavailable early in a pandemic due to limited diagnostics, or costly to obtain in low-resource settings. As cross-correlation values were highest for the preceding week (Figure 3), only search terms from that week were included in WSO model. The coefficients are presented in Table 4, for the model.
|
Variable |
Estimate |
Std. Error |
Pr(>|t|) |
|
Intercept |
-1336474 |
163324 |
2.39E-13 |
|
covidt-1 |
33447 |
2343 |
< 2e-16 |
|
COVID-19t-1 |
-30682 |
3285 |
3.91E-16 |
|
fevert-1 |
45812 |
4879 |
2.98E-16 |
Table 4 Web-search-only model (WSO) coefficients
Where 𝛽1, 𝛽2 and 𝛽3 represents the model coefficients for covid, COVID-19, and fever respectively. As with the previous models, the 1-step-ahead forecast had a smaller MSE than the 2-step-ahead forecast (112.486 × 10⁹ vs. 125.386 × 10⁹), but both were substantially larger than those of the VAR and TFM models. Although this model clearly overestimated case counts and peak values, it remained useful for predicting the timing of epidemic onset, peak, and decline. Figure 7 shows the model performance on the test dataset (Table 5, Figure 7).
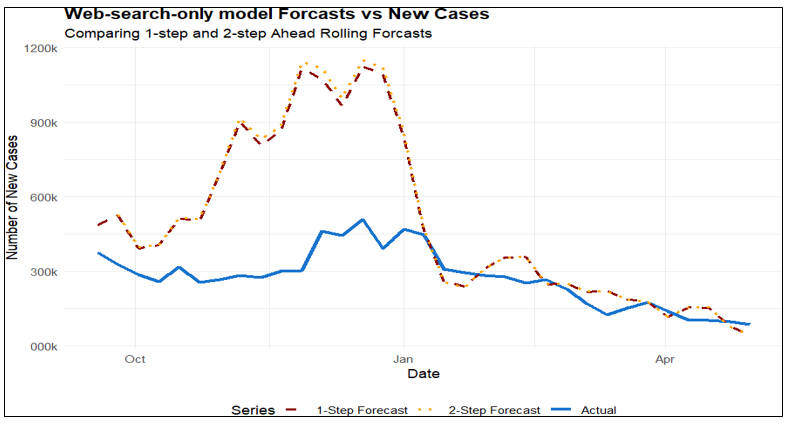
Figure 7 Web-search-only model: time series plot of new cases with 1-step-ahead and 2-step- ahead rolling forecasts.
|
Model |
AIC |
BIC |
MSE |
|
Vector auto regression |
4348.65 |
4388.8 |
1-step ahead |
|
6.573 x109 |
|||
|
2-steps ahead |
|||
|
13.075 x109 |
|||
|
Transfer function model |
3560.68 |
3572.21 |
1-step ahead |
|
9.941 x109 |
|||
|
2-steps ahead |
|||
|
25.913 x109 |
|||
|
Web-search only model |
3834.589 |
3849.003 |
1-step ahead |
|
112.486 x109 |
|||
|
2-steps ahead |
|||
|
125.386 x109 |
Table 5 Comparing models performance
Evaluation criteria for each model are summarized in Table 5 to compare the three models.
Our analysis demonstrated that parsimonious search terms such as covid exhibited stronger correlations with case counts than official terms like COVID-19; similar confounding effects were noted for terms such as flu.12 Between the two primary models, the VAR approach provided more accurate predictions of new cases and epidemic peaks (forecasted peak 604,896 vs. observed 508,009), compared with the less precise transfer function model (forecasted 645,808 vs. observed 508,009). The web-search-only model predicted the timing of epidemic onset, peak, and decline but greatly overestimated case counts, although underreporting in traditional surveillance systems may partly account for this discrepancy.8
In this study, we showed that COVID-19 cases can be accurately predicted using a combination of Google search trends for the term covid from the preceding week and new case trends from previous weeks. Both search and case trends demonstrated higher predictive accuracy with a one-week lag. Although the web-search-only model was limited to predicting the timing of epidemic onset, peak, and decline—and substantially overestimated case counts—we found it to be useful in underserved settings where weekly COVID-19 reporting was unavailable. This approach might allow researchers to glean useful information about outbreak dynamics even when case data are limited. Furthermore, comparison of correlations before and after attainment of 70% vaccination coverage showed that associations between search trends and new cases strengthened once the majority of the population had been vaccinated. This finding by itself suggests that once vaccination coverage surpassed 70%, Google search trends became more reliable predictors of new COVID-19 cases, likely reflecting reduced variability in disease dynamics and more stable population-level behavior.
This study provides valuable insights into predicting COVID-19 cases using prior-week search trends and case counts, but several limitations should be noted. For newly emerging diseases, it may take several weeks before reliable correlations between search activity and case incidence can be identified, limiting the model’s utility in the earliest stages of an outbreak.
Initial news coverage can also drive search behavior unrelated to actual symptoms, introducing bias. Furthermore, overlapping symptoms with other conditions, such as influenza, can confound search-based predictions (e.g., fever searches may reflect either flu or COVID-19).
Despite these challenges, the models evaluated here offer a useful starting point for anticipating case trajectories and highlight the broader potential of Syndromic surveillance. In conclusion, our findings reinforce the role of digital data streams as complementary tools to traditional surveillance, particularly in rapidly evolving public health crises. Beyond COVID-19, these approaches could be adapted to monitor future infectious disease outbreaks, helping to bridge gaps in early detection and response. Continued refinement of such models, paired with robust validation against clinical and epidemiological data, will be essential to ensure their reliability and to maximize their value for public health decision-making.
We would like to thank Dr. Rajabather Velu for his valuable suggestions on an earlier version of this study.
The authors declare there are no conflicts of interest.

©2025 Mohamed, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.
2 7