Advances in
eISSN: 2573-2862


Review Article Volume 3 Issue 2
Department of Veterinary Pathology, University of Maiduguri, Nigeria
Correspondence: Igbokwe IO, Department of Veterinary Pathology, University of Maiduguri, PMB 1069, Maiduguri, Nigeria
Received: April 24, 2017 | Published: March 7, 2018
Citation: Igbokwe IO. Evolving anti-disease strategies from biochemical pathogenesis of African trypanosomiasis. Adv Cytol Pathol. 2018;3(2):33-39. DOI: 10.15406/acp.2018.03.00048
African trypanosomes are transmitted biologically by tsetse flies (Glossina spp) to humans and animals to cause African trypanosomiasis or trypanosomosis (sleeping sickness or Ngana). The pathology associated with the disease includes blood dyscrasia (anemia, leucopenia, thrombocytopenia), organ damage and inflammation. The infected host’s metabolism is altered in response to the disease. The biochemical configuration and functions of the cell membrane, mitochondria, and cytosolic components are deranged by the infection. Cells of the infected host respond biochemically by producing chemical signals which regulate biochemical pathways, influence inflammatory responses and modify structural components. Several biochemical tools have been used to elucidate the biomolecules expressed in the disease. These biomolecules are also markers for diagnostic purposes and pathophysiological evaluations. The technologies involved in the biochemical studies are expanding from basic techniques of colorimetry, spectophotometry, and chromatography to sophisticated mass spectrometry. There are new trends employing systematics such as genomics, proteomics and metabolomics (metabonomics). The goal is to identify molecules and biochemical loci which can provide avenue for anti-disease strategies needed to treat and control the disease.
African trypanosomiasis is an important protozoan disease of humans and animals in many sub-saharan African countries where the disease is transmitted biologically by tsetseflies (Glossina spp) or mechanically by biting flies (Tabanids and Stomoxys). The pathogenic trypanosomes have specific hosts (Figure 1). The diseases caused by the trypanosomes are characterized by anemia, leucopenia, thrombocytopenia, inflammation of organs, tissue degeneration, apoptosis and necrosis,1,2 Cellular pathology caused by the disease is explored through researches using tools in biochemistry and molecular biology which examine biofluids, biomembranes, cytosolic compositions and capture molecular fingerprints associated with the disease processes. The review is aimed at briefly highlighting some of the biochemical elucidations of the pathogenesis of African trypanosomiasis from which anti-disease strategies would evolve.
Genomics in trypanosomiasis research
Molecular biologists have sequenced the trypanosome genome.3?6 The entire genome contains about 10,000 genes and 10% of them are variable surface glycoprotein (VSG) genes. The VSG genes provide a molecular basis for antigenic variation in trypanosomes during infections. Only one VSG gene is expressed at any particular time while others are transcriptionally silent. Antigenic variation enables the parasite to evade the host�s immune defence and inundate the host with a heavy antigenemia as the surface coat is discarded regularly. The dynamics VSG gene expression is expected to expose the conserved portions of the VSG proteome that is yet to be fully understood, and knowledge of it may be the basis for vaccine development against trypanosomes.7 Furthermore, trypanosome genomics may be a useful means of refining parasite identification, comparing species and life cycle stages, targeting gene locations for new drug development and improving understanding of drug intervention and resistance development. Recently, the pathogenicity of trypanosomes was linked to multiple genes, which implied that the differences in disease processes caused by various trypanosome species might have genetically-based incitement from the parasites.8 Variation in susceptibility of infected hosts to the pathogenic effects of trypanosomes has a connection with the host�s genetic resources. Genetic resistance to African trypanosomiasis is termed trypanotolerance and it occurs in some breeds of livestock, species of wildlife, strains of laboratory animals and humans.9,10 Trypanotolerance is an inheritable trait with genetic parameters of indicators such as control of parasitaemia and limiting of pathological effects.11,12 In cattle breeds, trypanotolerance arises from improved parasite control and haemtopoietic tissue genotype which improves control over development of anaemia.13,14 The capacity to limit anaemia development is more important for survival and productivity in cattle than parasite control.15 Therefore, understanding the genetic and transcriptional factors involved in trypanotolerance is relevant to the control of the disease9 and advances in host genetics in relation to infection are expected to help in the evolution of anti-disease strategies.12 Whole transcriptome analysis has revealed the gene variations related to the trypanotolerance trait in cattle.16 Trypanotolerance was associated with increases, at the early phase, in transcripts for genes encoding proinflammatory cytokine mediators during infection of N�Dama cattle.17 Temporal peripheral blood mononuclear cell gene expression in response to trypanosome infection was also recognised to be contributory to trypanotolerance in N�Dama cattle.18 There were differential gene expressions between trypanotolerant and trypanosuceptible breeds which underscored the genetic basis of the susceptibility traits.18 Serial analysis of gene expression (SAGE) from peripheral blood mononuclear cells led to the emergence of bioinformatic identification of upregulated and downregulated genes involved in trypanotolerance in cattle.19 Tumour necrosis factor gene was reported to play a role in resistance to trypanosome infection in mice.20 Higher serum amyloid A expression levels may support trypanotolerance in cattle.21 Host genetic deficiencies of complement lectin pathway factors contribute to infection susceptibility and disease progression.22
The genetic approach to understand the identified mechanisms of natural resistance to trypanosomiasis23 is supposed to provide insights on strategies for breeding selections so that the animal production system can insure the purity of genes of trypanotolerant breeds in livestock populations, and hence control the scourge of animal trypanosomiasis in endemic regions.24 It is noteworthy that introgression of genes of trypanosusceptible breeds into West African dwarf goat populations is occurring leading to loss of their trypanotolerance.25 Therefore, there is need to explore the host genome for the genetic architecture of the trypanotolerance trait and identify candidate genes to be recognised as selection signatures.26 In line with this proposition, several attempts have been made to characterise the candidate genes for trypanotolerance in cattle and ascertain whether the genes are under positive selection.27?29 Further details on the genetic basis of trypanotolerance across various species are available in a recent review by Yaro et al.,24 and the limitations of genetic studies with regard to their application in genomic selection are also reviewed by Meuwissen et al.,30 offering insights into improved genomic selection in animal breeding.
Biochemical interaction of trypanosomes with infected host
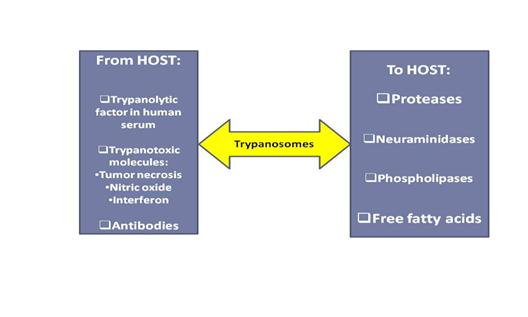
Trypanosomes, in body fluids, consume the host�s supplies (Figure 2) of carbohydrates, proteins, lipids and some micronutrients as their source of nutrients and release metabolites such as pyruvate, phenylpyruvate and tryptophol into the blood circulation as reviewed by Igbokwe.1 The trypanosome metabolites may cause metabolic acidosis and inhibit tricarboxylic acid cycle and glycogenesis in the infected hosts.1,31 The parasites (Figure 3) also produce proteases, neuraminadases, phospholipases, lipopolysaccharides and free fatty acids which contribute to the pathogenic mechanisms of the disease.1 Antigens are released from trypanosomes to stimulate the immune system of the host, which produce biomolecules such as immunoglobulins, lymphokines, nitric oxide and free radicals for defence against the infection.1 A trypanolytic factor in human serum, apoL-I, in high density lipoprotein is lethal for T. brucei.32 Trypanotoxic molecules (tumor necrosis factor, nitric oxide and interferon) produced by macrophages kill the parasites and antibody-opsonised trypanosomes are phagocytosed.33
Biochemistry of erythrocyte abnormalities
Erythrophagocytosis, after molecular alteration of erythrocyte membrane surface (Figure 4), in the macrophage phagocytic system (MPS) is the hallmark of trypanosomiasis.1 Trypanosome proteases and neuraminidases cleave surface sialic acids from surface membranes of erythrocytes and render them susceptible to phagocytosis.1,34,35 Also, oxidative stress in trypanosome infections has implications on erythrocyte membrane stability and fluidity. Erythrocytes with altered membranes following peroxidative injuries would be phagocytosed or fragmented and lysed intravascularly.1,34,?41 The transfer of VSG from trypanosomes to erythrocyte membranes and the anchorage of immunologically derived molecules on erythrocyte membranes have been shown to lead to erythrocyte clearance by MPS.42 Perturbation of membrane function has been suggested by the report of decreased erythrocyte potassium and increased erythrocyte calcium ion contents in trypanosome infections in association with decreased osmotic resistance.43 It was presumed that the functions of NaK-ATPase and CaMg-ATPase may be affected in the infection,1 because these membrane-associated enzymes were reduced in activities during infection in the brain, kidney and testis.44?46 Mijares et al.,47 confirmed in their report that erythrocyte Ca-ATPase activity decreased in Trypanosoma evansi infection. Trypanotolerance has been associated with higher erythrocyte levels of zinc and manganese,48 increased erythrocyte sialic acid content,49 differential erythrocyte sialic acid composition50 and erythrocyte surface sialic acid moieties characterised by more O-acetyl than glycolyl groups.51
Biochemistry of dyserythropoiesis
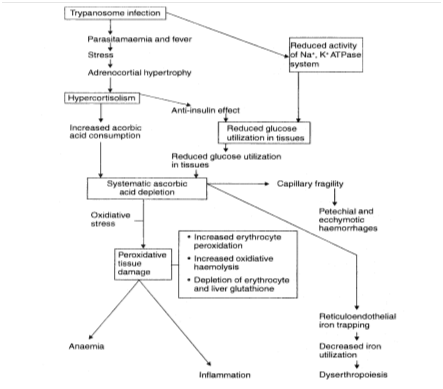
The capacity of an infected host to resist anaemia, and respond to it through enhanced erythropoiesis, are valuable traits in trypanotolerance. More than two decades ago, there was indirect evidence that erythropoietin activity might be reduced in the plasma of trypanosome-infected animals.52 The finding was followed with a hypothesis that trypanosome neuraminidase could destroy the in vivo biologic activity of erythropoietin (EPO), a glycosylated protein, by cleaving the terminal sialic acid residues53 and this was subsequently supported by others.34 Later, it was reported that EPO activity increased in infected calves that had high erythropoietic potential, but the EPO had impaired activity because of inhibitory factors in the plasma, thus supporting our earlier findings.54,55 Suliman et al.,56 reported that the levels of EPO and receptors of EPO in infected Boran cattle were inadequate for the degree of anaemia during trypanosome infection. Downregulation of EPO receptor expression seems to impact the proliferation, differentiation, and maturation of the erythroid precursors and leads to inadequate erythropoiesis in the infected animals.57 The survival rates of infected mice were dramatically improved by treatment with recombinant human erythropoietin,57 which is relevant to the report that increased erythropoietic potential and ability to prevent anemia are related to trypanotolerance.15 Trypanotolerance was associated with higher levels of EPO receptor transcripts in the bone marrow.56 Strong immune response (Type-1) is mounted during the early stages of infection and it is associated with the classical hyperactivation of macrophages which are engaged in Erythrophagocytosis.1,33 The engulfed erythrocytes release haemoglobin from which iron (Fe2+) is extracted in the phagosome and moved into the cytosol in the ferrous state to be converted to ferritin (Fe3+) for storage. Extracellular exportation of ferrous iron occurs on demand via ferroportin-1 (FPN-1). The extracellular Fe2+, upon conversion to ferric iron (Fe3+) through ceruloplasmin (CP), is bound to transferrin (Tf) and transported, mainly to the bone marrow to be used in erythropoiesis. In trypanosome-infected mice, there were increases in gene expressions of FPN-1, Tf and CP only in the acute phase but not in the chronic phase, suggesting that export of iron was hampered in the chronic phase of infection with indication of iron sequestration, as evidenced by increased ferritin expression, enhanced uptake of erythrocytes and iron-containing compounds.58 Depletion of ascorbic acid pool of the host during the infection was postulated as a possible cause of the iron sequestration (Figure 5), since ascorbic acid is required to mobilize iron from the ferric state of the stored iron to the ferrous state for exportation to the extracellular compartment en route to the bone marrow31 and supplementation of the vitamin may eliminate the iron blockade and allow enhanced erythropoiesis after correction of any ascorbic acid deficit as already report in trypanosomiasis.59 The depression of serum levels of some free amino acids occur in trypanosome infections and may be responsible for the reduction in the rate of synthesis of the globin moiety of haemoglobin as previously reviewed.31 Globin synthesis precedes heme synthesis and is nearly complete by the time haemoglobin synthesis begins. Thus, reduced availability of required amino acids for globin synthesis may also slow down erythropoiesis.
Oxidative stress in trypanosome infections
Oxidative stress is an imbalance between free radical generating and scavenging activities, resulting in the accumulation of oxidation products which are capable of causing tissue damage. A free radical is highly reactive and can be produced from metal or enzyme catalyzed redox reactions of cells. Because bloodstream forms lack catalase and GSH peroxidase activities, hydrogen peroxide accumulates in the trypanosome to a concentration of about 70 �M.60,61 Therefore, hydrogen peroxide and free radicals may be discharged into the extracellular fluid by the parasite in infected animals. Activated macrophages are capable of increasing their oxygen uptake after which they release large amounts of superoxide anions into the extracellular fluid.62 The generation of free radicals is mediated by pro-inflammatory cytokines such as tumour necrosis factor alpha (TNF-a), gamma interferon (IFN-g), interleukin1beta (IL-1B) and IL-6 produced by activated macrophages.33 Malondialdehyde (MDA) is the most abundant biomarker produced as a consequence of lipid peroxidation after free radical attack of cellular lipids.63 The accumulation of MDA in tissues or biological fluids is indicative of the extent of free radical generation, oxidative stress and tissue damage.64 Reports have shown that trypanosome infections increase MDA production in the host as an indication of oxidative damage.37,65
Methemoglobin (MHb) is the oxidation product of haemoglobin which occurs when ferrous iron in hemoglobin is oxidized to the ferric state and may aggravate oxidative damage to erythrocytes. Increased MHb in T. evansi-infected camels provide further evidence of erythrocytic oxidation and free radical generation in the erythrocytes during infection.37 Nitric oxide was generated in excessive amounts in T. brucei, T. b. rhodosiense T. evansi-infected individuals.37,66,67 Nitric oxide may react with superoxide anions to produce peroxynitrite anion (PNA), or by the Fenton reaction to produce hydroxyl radical.68 Both PNA and hydroxyl radicals are the most potent oxidizing agents which can initiate lipid peroxidation in the cell membranes causing cellular damage.63 Furthermore, nitric oxide can cross the erythrocyte membrane and react with SH group resulting in nitrosyl-haemoglobin, which slowly oxidizes to MHb in the presence of oxygen.69 Direct correlation of nitric oxide production with the development of anaemia in T. brucei-infected mice was reported, and the treatment with nitric oxide blockers led to a significant reduction of the anaemia.67 Positive correlations of enhanced nitric oxide production with erythrocyte MDA generation and MHb formation were reported as evidence of the involvement of nitric oxide production in erythrocytic oxidation during T. evansi infection while the negative correlation of nitric oxide, erythrocyte MDA and MHb formation to packed cell volume indicated that the enhanced oxidation of the erythrocytes contributed to the progression of the anaemia in chronic T. evansi-infected in camels.37 The antioxidant systems involved in the protection against free radicals are superoxide dismutase (SOD), GSH-peroxidase, catalase (CAT), glutathione (GSH), antioxidant vitamins A, E and C. When the antioxidant systems are challenged by free radical generating events during the disease process, the activities of the antioxidant enzymes are affected and the antioxidant molecules are depleted. The activity of erythrocyte GSH peroxidase was increased in trypanosome-infected mice,40 whereas erythrocyte SOD activity decreased in T. evansi-infected camels.37 Most importantly, antioxidant GSH and vitamins A and C are depleted in these infections.38,59,70 The assessment of the antioxidant deficit of erythrocytes in trypanosome infections was also done by in vitro peroxidation test where the cells were challenged with peroxide load and found to have increased susceptibility.36 Increased susceptibility of erythrocytes to oxidative hemolysis was also an indication of the decline in antioxidant capacity of erythrocytes during infection.71
New approach to metabolic studies in infected hosts
Cellular homeostasis depends on cell membrane integrity, respiration and energy generation. Any deviation means a tendency towards cellular degeneration, a reversible cellular state or cell death, a terminal state leading to cellular disintegration. Indications that trypanosome infections affect cell metabolism are being discovered and areas of science known as proteomics and metabonomics are deployed to improve our understanding. The central issues on metabolic dysfunctions during infections include the increases in catabolic processes, downregulation of energy generating pathways, depletion of energy molecules with depressed intracellular glucose delivery and loss of capacity to endogenously generate antioxidant capacity.31,72 With proteomic studies, the protein configurations of the altered state are studied with better efficiency and these protein expressions from the parasite and host cells provide opportunities in drug development.73 Also, metabolic profiling enables us track the systemic weakness that develop with the progression of the disease in order to present a global perspective to the pathophysiology of the disease and give insights into most metabolic dysfunctions.74
Anti-disease strategies against trypanosomiasis
Strategies conceived to reduce or eliminate the burden of the disease in the parasite-host interaction, would be expected to mitigate paradigmatic parasitism, where parasite population increases and the host loses fitness from organic lesions associated with systemic dysfunctions. In this context, anti-disease strategies against African trypanosomiasis, in the broadest sense, may include: a) prevention of infection by breaking trypanosome transmission through avoiding contact with vectors by using odour-baited traps and screens to remove vectors from the vicinity of the host,75 deploying synthetic or natural animal-derived vector repellents on the host76,77 and/or elimination of vectors in the environment by aerial insecticide sprays78?81 and integrated vector control using sterile insect technique;82,83 b) reduction of the virulence or attenuation of trypanosomes by genomic or transcriptional manipulation during propagation in the vector or host84,85; c) development of a vaccine and vaccination to prevent disease through protective immunity86; d) enhancing natural innate resistance or immunity17,20,87?89 that controls parasitaemia by genomic selection of the host population;26,30 e) reduction of host immunopathological responses responsible for pathophysiologic conditions associated with the disease;33,90 f) interventions to ameliorate or abet somatic dysregulations and dysfunctions;91?94 g) clearance of parasitaemia by effective chemotherapy.95,96 Antoine-Moussiaux et al.,97 considered, in a review, that anti-disease strategies could be targeted at treatments which did not eliminate the trypanosomes, but prevented the invasion of the brain by maintenance of blood brain barrier, neutralized the toxic products of trypanosomes, modulated the immune processes towards regulation of inflammation, reduction of effect of tumour necrosis factor, amplification of TGF-beta 1 and prevention of apoptosis of lymphocytes. Therefore, the anti-disease strategies propose to promote disease tolerance which is an ability of the host to limit the health consequences of the infective pathogen in spite of burden of infection.98,99
The strategies directed against disease-producing pathophysiological processes to engender trypanotolerance or resistance to the disease require continuous attention in research, because several published results as reviewed here, so far, have provided insights as to the themes for re-evaluation and re-emphasis. It is imperative to keep this direction of research because of the failure of control of measures. Complete elimination of vectors has not been feasible in most endemic environments and multi-drug resistance has made chemotherapy less effective, causing relapse of patent infections in treated cases. Sometimes, diagnostic techniques may not be able to detect very low parasitemia and latent infections when the parasites are sequestered in the extravascular compartments. Trypanosomes in infected hosts migrate to privileged extravascular sites which may be adipose tissues, dermis and subcutis of the skin, where they localize and serve as sources for subsequent recrudescence of active patent infection.100?102 Therapeutic agents need to be capable of reaching tissues with sequestered parasites which are shielded from drug effects and in the event of failure to do so, the infected hosts has to recruit endogenous anti-disease strategies to control the infection or be supported exogenously with anti-disease interventions. Even in untreated cases, an argument is tenable for strategies investing in disease mitigation of the infected hosts. A valid approach is to evolve the anti-disease strategies from the understanding of the biochemical abnormalities produced in the development of the disease so that interventions could support the host in the averting or remedying the biochemical lesions.
None.
None.

©2018 Igbokwe. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.