MOJ
eISSN: 2373-4442


Mini Review Volume 6 Issue 3
1Department of Bioinformatics and Functional Genomics, Institute of Cytology RAS, Russia
2Latvian Biomedical Research and Study Centre, Latvia
Correspondence: OV Anatskaya, Department of Bioinformatics and Functional Genomics, Institute of Cytology RAS, St Petersburg, Russia
Received: January 01, 1971 | Published: June 7, 2018
Citation: Anatskaya OV, Erenpreisa JA, Salmina KA, et al. Polyploidy activates biological pathways related to morphogenesis in mammalian tissues. MOJ Immunol. 2018;6(3):90-93. DOI: 10.15406/moji.2018.06.00200
Background: Polyploid cells exist in all mammalian tissues where they were searched. In physiological conditions polyploidy is a part of development and differentiation programs. In stressful condition it may promote pathological processes, including carcinogenesis.
Aim and Methods: Currently, the impact of polyploidy on morphogenetic processes remains almost unstudied. To address this issue, we developed an integrative bioinformatic approach consisting in a pairwise cross species transcriptome comparison of homologous mammalian tissues with various degrees of polyploidy (Haman and mouse heart and liver).
Results: Our data indicated that polyploidy activates gene modules of embryogenesis and impairs gene modules of differentiation. Polyploidy activated genes were enriched (i.e., were presented above the random level) in GO biological processes and in KEGG pathways related to organism development, morphogenesis, and stem cell biology (including the pathways of Hippo, Pi3K, WNT, Hedgehog, and TGF-ß signaling), whereas polyploidy inhibited genes were enriched in gene modules related to differentiation. The structure and composition of the protein interaction networks constructed for polyploidy regulated genes confirmed the results of gene module analysis.
Conclusion: Thus, our results demonstrate that somatic cell polyploidy can regulate the modules of organism development by increasing their activity.
Keywords: polyploidy, cross-species comparison, transcriptome, morphogenesis, human, mouse
Polyploid cells contain several copies of diploid genomes. These cells were found in all mammalian tissues where they were searched. In physiological conditions, genome accumulation is a part of development and differentiation programs. Polyploid cells present in heart, skin epithelium, placenta, liver, brain, and blood.1 In stressful condition polyploidy accompanies and stimulates pathological processes. Thus, the increase of polyploidy was documented in hypertensive heart, atherosclerotic vessels, cirrhotic liver and cancer.2 Despite ubiquitous distribution, the effect of polyploidy on gene expression remains not completely understood. Increasing evidence suggests that cell polyploidy may epigenetically alter gene expression and miRNA gene transcription in plants as well as in animals.3,4 miRNA which are short (20-22 nucleotide long) noncoding RNA reported to play various roles in human, plant and in some viruses.5,6 The impact of polyploidy on the transcriptome was investigated in hepatocytes, megakaryocytes, vascular epithelial cells, and placenta. Investigation of the differences in expression using the conservative double threshold led investigators to the conclusion that among several thousand genes not a single one is affected by polyploidy (as in the case of hepatocytes) or only several dozen genes are reactive.7 Moreover, for each cell type, this small set of genes turned out to be different. For instance, in the cells of vascular epithelium, several genes associated with the immune system were suppressed;8 in megakaryocytes, activation of tissue specific genes was registered; and, in decidual cells, activation of nuclear genes encoding mitochondrial proteins was detected.9 These differences suggest that the full effect of somatic polyploidy (if it exists) has not been identified yet. The absence of consensus is not surprising because mmaintaining gene dosage balance, polyploidy has little impact on separate gene activity. The weakness of ploidy associated effects make it difficult to identify common gene expression signature. At the same time, modifying activity of thousands of genes at a time, polyploidy participates in coordination of postnatal development, developmental programming, differentiation and pathogenesis of many diseases, including cardiovascular diseases and cancer.2,7,10–12
Currently, the impact of polyploidy on morphogenetic processes remains almost unstudied. To address this issue, we developed an integrative bioinformatics method aimed at accessing weak effects of polyploidy on the activity of genes by implementing a pairwise cross-transcriptome analysis of mammalian tissues with varying degrees of ploidy.13–16 The aim of the present study was to investigate the effects of polyploidy on activity of gene modules related to development, embryogenesis, morphogenesis and differentiation by the method of extensive cross species transcriptome comparison.
The basis of the applied bioinformatic approach consists in simultaneous pairwise comparison of transcriptome activity across tissues and between species in homologous organs with different degrees of polyploidy, for example, in the heart and liver of humans and mice. Our previous studies have shown that human heart manifests high ploidy (four to eight genomes per cardiomyocyte nucleus), while human liver is predominantly diploid (about two genomes per hepatocyte nucleus). Mice, on the other hand, have diploid heart (about two genomes per cardiomyocyte nucleus) and a highly polyploid liver (four to eight genomes in the nuclei of hepatocytes).13 Further, for simplicity, we will use the term “polyploid organs” to denote the composition of mainly polyploid cells, such as in human heart and mouse liver; and “diploid organs” to refer to the consistency of mainly diploid cells, such as in mouse heart and human liver. The source of data for these studies was the BioGPS database17 containing full transcriptome data obtained using oligonucleotide microarrays. We selected only well characterized genes, i.e., the ones presented in the NCBI Gene database.18 Furthermore, genes were required to be homologous (orthologous) in human and mouse.
Complete human and mouse proteomes were obtained from the RefSeq database.19 We employed the highly sensitive Smith–Waterman algorithm realized in the Search program within the Fasta software package20 to perform a whole genome comparison for each human protein versus all mouse proteins (and, vice versa, each protein of the mouse versus all human proteins). After that, we selected pairs of genes with the greatest statistical significance for encoded proteins in both cross comparisons (reciprocal best hit), which can be considered orthologous. After selecting pairs of orthologous genes, we carried out analysis of changes in expression in each pair. We considered that genes changed activity depending on the ploidy if gene expression changed in the same direction in both reciprocal comparisons in line with the difference in the degree of tissue ploidy (human heart → mouse heart; mouse liver → human liver). Genes that met all these criteria have been divided into activated and inhibited ones and analyzed for enrichment in biological processes in the Gene Ontology (GO) database21 using the approach developed in.22 As a positive enrichment, we considered a statistically significant excess over a randomly expected level. In particular, we used the average change in expression levels for both cross comparisons (human heart → mouse heart; liver of the mouse → human liver).
Gene module enrichment analysis
Functional groups and pathways enriched for differently expressed genes were identified based on GO and KEGG pathway analysis as described previously14,22 Protein interaction networks for gene products that play a role in the GO biological processes regulating stem cell biology were constructed using the STRING server.23 The enrichment significance of GO biological processes and KEGG pathways in protein products from obtained networks were also determined using the STRING database server.23
To find out how polyploidy affects gene modules related to development, we applied pairwise cross species transcriptome comparison for human and mouse heart and liver.16,22 It is well established that human and mouse have the inverse patterns of polyploidization in cardiomyocytes vs. hepatocytes,22 which enables the removal of the effect of species and tissue-specificity on the profile of gene expression while comparing homologous tissues contrasted in ploidy levels. Gene activity was considered dependent on ploidy if the ratio of the amount of mRNA transcript in two pairwise comparisons for polyploid and diploid organs (human heart/mouse heart and mouse liver/human was greater than 2.0 fold.
The comparison revealed more than 700 genes with the increased and more than 500 genes with the decreased levels of mRNA in polyploid vs diploid tissues. Of a total list of GO categories that were significantly enriched in ploidy regulated genes, 89 contained in their titles terms “development”, “embryo”, “morphogenesis”, “differentiation” and “stem cell” and thus were of particular interest. Of 89 processes, 79 related to development and differentiation (Figure 1A) and 10–to stem cell proliferation and maintenance and contained differentiation”. (Figure 1B) GO processes coordinating development, morphogenesis, proliferation and stem cell maintenance were enriched in ploidy-induced genes. GO processes implicated in cell differentiation were enriched in ploidy inhibited genes (Figure 1A) (Figure 1B), confirming a causal link between ploidy and induction of fetal and morphogenetic programs. Also we found that ploidy activated genes were enriched in biological processes of Gene Ontology (GO) database and in pathways of KEGG database related to multipotency, including Hippo, Pi3K, WNT, Hedgehog, and TGF-ß signaling to a greater extent than genes suppressed by ploidy.
Currently, there are only a few studies evidencing that polyploidy activates developmental pathways. For instance, a positive correlation was established between hyperpoliploidy of cardiomyocytes and expression of fetal phenotype determined based on the induction of embryonic markers: beta-myosin heavy chain, factor HIF1α (hypoxia factor 1α), an increase in the number and size of the nucleoli, as well as increased proliferation potential.9–13 In accordance, our recent study indicated that polyploidy boosts Myc interacting genes and pathways of multi-and pluripotency.
Network of protein interactions for genes from modules contributing to stem cell biology related pathways
Genes regulating stem cell biology play a central role in coordination of developmental programs. Therefore, it was important to investigate features of their functional interactions. For this propose, we constructed a network of protein–protein interactions for genes that are involved in regulation of gene modules contributing to stem cell biology, i.e. modules containing in their titles term “Stem”. (Figure 1B) Overall, we identified 24 proteins with these properties. (Figure 2A) All of them belonged to GO biological processes indicated at Figure 1B.
To investigate in more detail biological functions of the 24 nodes (genes) of the network, we identified GO biological processes and KEGG pathways that are enriched in these genes using String server23 Modules and protein interaction networks can indicate a link between genes and biological functions, thus constitute a key step in connecting genotype and phenotype.24 Figure 2 shows modules with statistically significant enrichment in network forming genes. It is evident that the highest accuracy among GO processes (Figure 2A) is exhibited by the modules associated with development, differentiation, and proliferation of stem cells, as well as a pathway of epithelial–mesenchymal transition. It should be noted that almost all hub (node) proteins participate in several modules, which points to their plexus and synergism. Maximum activation of expression was registered for GO module of stem cell proliferation. This module consists of genes creating balance between early cell patterning and self renewal. Similarity in gene composition for processes of proliferation and differentiation can be explained by the fact that, for stem cells, these processes represent different stages of specialization. Development represents an earlier phase, and differentiation represents a later one.25
Assessment of KEGG pathways that are enriched for 24 network creating genes (Figure 2B) revealed statistically significant enrichment for pathways of Hippo, Hedgehog, WNT, and TGFβ signaling. There is ample evidence indicating that these pathways coordinate the regulation of developmental processes and cell pluripotency.26,27,28 Thus, predominance of activated genes compared to repressed genes, and preferential activation of important positive regulators (Myc, Taz, Kit, Wnt2, Wnt2b, Fgfr2) coinciding with a decrease in activity of a negative regulator of the Wnt pathway, Gsk3b,28,29 complement and support our data obtained with the analysis of GO processes. Reactivation of embryonic programs by the incidence of polyploidy has been previously recorded only in transformed cells and in cultures of aging and transformed fibroblasts.12,30,31 For this reason, the results obtained in our study for normal cells in living organisms can be considered as a first evidence of the existence of common features between normal and transformed cells.
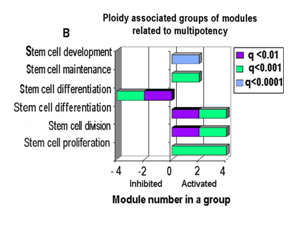
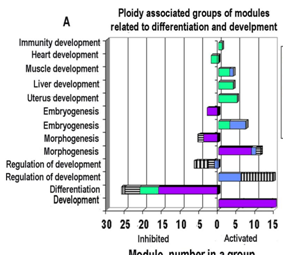

Figure 1 GO biological processes related to development, differentiation (A) and stem cell (B) that are enriched for ploidy regulated genes in simultaneous pairwise cross-species comparison of polyploid vs diploid organs (see methods, please): human heart vs mouse heart and mouse liver vs human liver comparisons. q– means p value with multiple comparison tests. SC–stem cells.
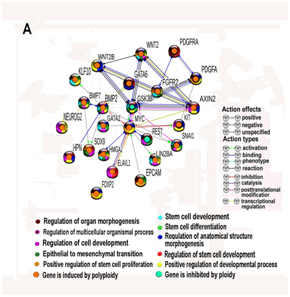
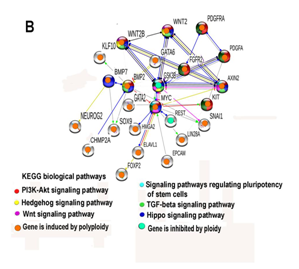
Figure 2 Gene module enrichment analysis of protein-protein interaction network composed by 24 ploidy associated genes that are involved in the regulation of stem cell biology. A-network enrichment for GO biological processes with q value not less than 10-11. B-network enrichment for KEGG pathways with q value not less than 10-4. Both networks constructed with stringency S<0.4. The networks were constructed using String server. All genes in the network show similar ploidy related changes in two simultaneous pairwise comparisons: human heart vs mouse heart and mouse liver vs human liver.
Somatic polyploidy is wide spread in normal mammalian tissues and increased at many pathological states, including carcinogenesis and degenerative disorders.32 At the same time, the physiological role of this phenomenon remains unclear. The initial hypothesis that genomic duplications are associated with terminal differentiation and aging has not been confirmed.7 Moreover, data obtained from studies on transformed cells demonstrate that polyploidy promotes activation of important markers of pluripotency.31 In this respect, information on the effect of somatic polyploidy on the transcriptional activity of modules of development and the structure of their regulatory networks obtained by our newly designed comparative transcriptome analysis approach will help to improve understanding of the role of polyploidy in normal physiology, transformation, and other pathological conditions.
The work was supported by Russian Science Foundation (Grant 14-50-00068).
The author declares there is no conflict of interest.

©2018 Anatskaya, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.