MOJ
eISSN: 2471-139X


Review Article Volume 3 Issue 1
Department of Cell Biology and Anatomy, Rosalind Franklin University of Medicine and Science, USA
Correspondence: Michael P Sarras, Department of Cell Biology and Anatomy, Rosalind Franklin University of Medicine and Science, 3333 Green Bay Road, North Chicago, IL 60064, USA
Received: December 09, 2016 | Published: January 20, 2017
Citation: Sarras MP. Laminins of hydra ECM: a comparative analysis of hydra and vertebrate laminins based on current genomic and ETS databases. MOJ Anat Physiol. 2017;3(1):11–15. DOI: 10.15406/mojap.2017.03.00078
Extracellular matrix (ECM) arose when the earliest metazoans with defined tissues diverged some 500 million years ago. The Cnidarians, which include Hydra, are one of these Taxa. Studies beginning in the 1980 used convergent biochemical, cellular, and molecular approaches to determine the structure/function relationships of Hydra ECM. Initial screening of cDNA expression libraries identified laminins as a prominent ECM component localized to the basal lamina of Hydra’s ECM. Those first laminins matched with the vertebrate laminin a (likely α1) and β1 chains. Structure/function studies determined that Hydra laminins were critical to the organism’s ECM structure and the ability of Hydra to regenerate. Since those studies, extensive genomic and EST studies have expanded our understanding of the full spectrum of laminin chains in Hydra (to a total of 6 vertebrate-like laminin chains) and this article will compare Hydra’s laminins to the vertebrate laminins which are known to be involved in embryogenesis, cell physiology, and numerous human diseases.
Keywords: hydra, extracellular matrix (ECM), laminin, vertebrates, genomic data bases, EST data bases
ECM, extracellular matrix; GF, growth factors; MMPs, matrix metalloproteinases; LAMA1-5, laminin alpha1-5 (α1-5); LAMB1-3, laminin beta1-3 (β1-3); LAMC1-3, laminin gamma 1-3 (γ1-3); Laminin-111, laminin heterotrimeric molecule containing a single α1, β1, and γ1 chain; LE domains, epidermal growth factor-like domain; G Domain, globular domain; EST, expressed sequence tags
The extracellular matrix (ECM) is a complex structural and signaling extracellular entity that has multiple cell physiological and developmental functions.1,2 These functions include:
Given the fundamental function of the ECM of metazoans, it is not surprising to find ECM components in ancient Taxa such as the Cnidaria that represents one of the earliest eukaryotic systems with a defined tissue organization.4 Cnidarians are composed of an outer ectoderm and inner endoderm with an intervening ECM. Based on molecular studies focused on understanding the components and function of various macromolecules of the ECM in Hydra, we now have a basic understanding of the role of ECM in an early divergent metazoan that arose some 500million years ago.5‒7
This review focuses on an important component of the ECM, namely laminin; the matrix molecule first cloned and characterized in the Cnidarian, Hydra. Its purpose is to show the highly conserved nature of ECM throughout the Metazoans by highlighting the structure/function relationships of an early laminin form. It does this by comparing this laminin to what we know of vertebrate laminins and our knowledge of the full spectrum of Hydra laminins based on current genomic8,9 (NCBI Hydra Genome Database: https://www.ncbi.nlm.nih.gov/genome/?term=Hydra) and EST data bases10 (NCBI Hydra EST Database: https://www.ncbi.nlm.nih.gov/nucest) studies. The review is organized to discuss
Initial cloning and characterization of a hydra laminin
From the standpoint of divergence of Taxa within the animal kingdom, Cnidarians are the earliest group of metazoans with a defined tissue organization (ectoderm and endoderm) and an ECM. A tree of Taxa divergence is shown in Figure 1. This tree of Taxa with a “Common Tool Kit” of ECM components indicates that Cnidaria, containing Hydra, arose some 500million years ago, thereby representing one of the earliest metazoan life forms with an ECM.5
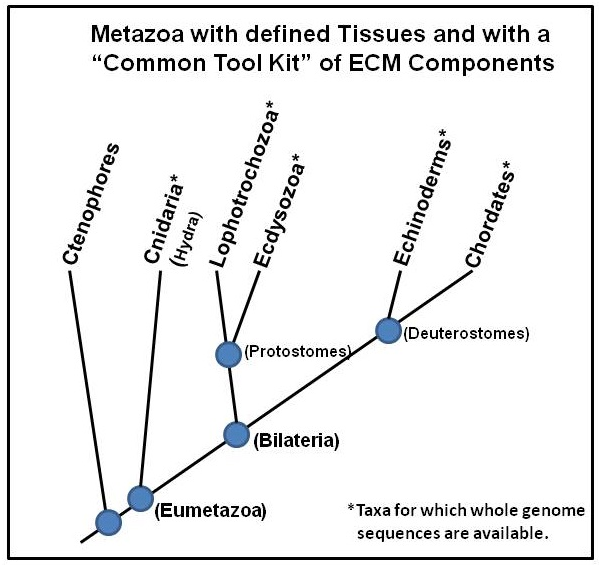
Figure 1 A hieratical tree of metazoans with a “Common Tool Kit” of ECM components. Those Taxa with an asterisk have complete genome data bases available. Cnidaria divergence dates back approximately 500 million years ago.
The basic structure of Hydra’s body wall is shown in Figure 2. As indicated, the entire body wall of Hydra (from the foot process, through the body column, and apically to the tip of head tentacles) is organized as an outer ectoderm and inner endoderm with an intervening ECM (Figure 2, left image). Based on molecular and cellular studies, Hydra’s ECM is itself organized as two basal lamina’s (adjacent to the two epithelial layers) with an intervening interstitial-like matrix composed of fibular collagens as shown in Figure 2, right image. Because of its simplified structure and high regenerative capacity, Hydra has been studied as a model developmental organism since the time of Trembley whose studies were published in 1744.7
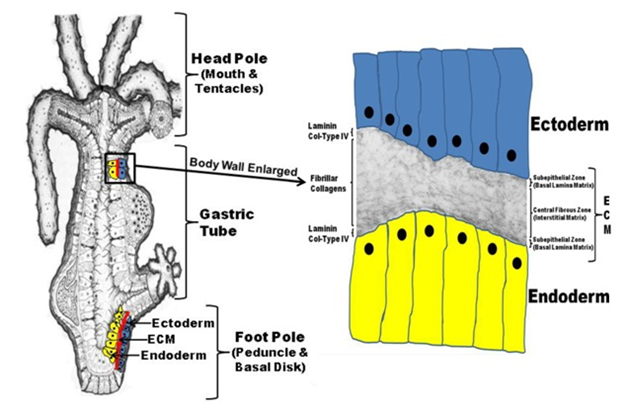
Figure 2 Hydra body plan formed from an epithelial bilayer with an intervening Extracellular matrix (ECM).
Hydra exists as a gastric tube with a mouth and several tentacles at the head pole and a peduncle and a basal disk at the foot pole (left diagram). The entire body wall of Hydra (from the tip of the tentacles to the basal disk) is organized as an epithelial bilayer with an intervening ECM along the entire longitudinal axis of the organism. Hydra’s ECM is structured as two subepithelial zones (i.e. basal lamina matrix) with an intervening central fibrous zone (i.e. interstitial matrix). As shown in the composite diagram to the right that utilizes a transmission electron micrograph of Hydra ECM interposed between a drawing of the two cell layers (ectoderm and endoderm), Hydra laminin and Type IV Collagen are localized to the two subepithelial zones of the matrix while Hydra fibrillar collagens (e.g. Hcol-I) are localized to the central fibrous zone or interstitial matrix.
Using a combination of biochemical, cellular, and molecular approaches, studies were initiated in the 1980’s to elucidate the structure and function of hydra ECM components.11 The work involved:
While a number of Hydra ECM components were identified in this way, for the purposes of this review we will focus on those studies concerning Hydra laminin, the first molecule to be clearly identified and functionally characterized. Laminin chains were identified using antibodies generated to purified Hydra ECM. Initial antibody screening of expression cDNA libraries isolated clones matching the laminin a chain (likely α1) and β1 chain of vertebrates.13,15 The complete ORF of the β1 chain was determined while only a partial sequence of the a chain (likely an α1 chain) was obtained. While Hydra laminin is localized to the two subepithelial zones (basal lamina) of Hydra ECM, it is synthesized exclusively by the endoderm, which means that the molecules have to diffuse through the mesoglea to reach the ectodermal layer.13,14 The location at the basal region of both cell layers (Subepithelial Zones of Hydra ECM) as shown in Figure 2 suggests that laminin is required for proper cell function and differentiation. During Hydra ECM regeneration, laminin secretion from the endoderm precedes the secretion of Hydra collagen-I that arises from the ectoderm16 and inhibition of laminin secretion through a RNA antisense technique will block collagen secretion;17 indicating cross-talk between the two layers. Earlier studies have already established that antibodies to Hydra laminin will block Hydra morphogenesis14 and other ECM-related processes such as cell migration.18 mRNA expression studies found that laminin is up-regulated during regeneration and in situ expression studies found this up-regulation to be associated with the base of head and foot regeneration regions.17 Potential cell surface binding domains were later identified in the laminin β1 chain along with cell binding proteins that had affinity for these binding domains.13,19 As indicated, laminin expression was essential for Hydra morphogenesis and regeneration processes based on a number of functional studies.17
These studies indicated that ECM components, such as laminin, formed early during the divergence of early metazoan groups (e.g. Cnidaria) and these molecules were essential for Hydra cell physiology and developmental/regenerative processes.
Composition of vertebrate laminins
Vertebrate laminins1,2 are heterotrimeric cross-linked glycoproteins that bind to other matrix components in order to form the complete basement membrane network typically associated with epithelial cells. Laminins predominantly link with collagen IV, nidogens, agrin, and perlecan to form the ECM basement membrane (termed the basal laminin at the level of transmission electron microscopy).20 Laminins are composed of three chains, named the alpha (a), beta (b), and gamma (g) chains, that are coded by separate genes. Genomic and biochemical studies with vertebrate systems have identified five a chains, three b chains, and three g chains. These chains are assigned the symbols; LAMA1-5, LAMB1-3, and LAMC1-3. In vivo, we find 16abg laminin trimers that exist in nature. The reduced number of laminin trimer chain combinations (as compared to the theoretical maximal number of combinations) is due to the biochemical properties of the chains and the distribution of laminins among the various tissues of the body. The most studied heterotrimeric laminin is composed of a α1, β1, and γ1 chain and is termed laminin-111. A diagram of laminin-111 is depicted in Figure 3.

Figure 3 A diagram of the trimeric laminin-111 molecule. Laminin-111 is composed of a single α1, β1, and γ1 chains. A description of the various regions of laminin is discussed in the text.
As discussed, since the original identification of the first laminin in vertebrates, a total of 11 laminin chains have been identified and 6 of these chains have analogs in Hydra (see Section 3 & Table 1). The biochemistry of laminin chains has been well studied.1,2 All laminin chains have common structural motifs that include:
Laminins have a critical role during many systemic processes such as embryogenesis,20 wound healing,21,22 and angiogenesis. In addition, laminins are central to many human diseases such as, muscular dystrophy type-1A,23 Pierson syndrome, epidermolysis bullosa,24‒26 laryngo-onycho-cutaneous syndrome,27 and tumorigenesis.28,29
Hydra Laminins Identified from Genomic and EST Analyses |
|||
Genomic Analyses |
EST Analyses |
||
Alpha chains |
Gene ID # |
Alpha chains |
Accession #: |
Laminin α 1 |
100197510 |
Laminin α 1 |
DN815114.2 |
Laminin α 2 |
101235032 |
||
Laminin α 3 |
101241783 |
||
Beta Chain |
Gene ID # |
Beta Chain |
Accession #: |
Laminin β1 |
100202556 |
||
Laminin β2 |
101236303 |
||
Gamma chain |
Gene ID # |
Gamma chain |
Accession #: |
Laminin ϒ1 |
100198511 |
Laminin ϒ1 |
DT613962.1 |
Table 1 Provides a list all identified Hydra laminins
Composition of hydra laminins based on the most current NCBI data bases derived from various genomic and EST studies
Since the cloning and functional characterization of the Hydra laminin β1 chain and the Hydra laminin a chain (likely α1), no other articles have been published that focus on the detailed characterization of the other Hydra laminins. However, with the development of full genomic sequence data bases8,9 (NCBI Hydra Genome Database: https://www.ncbi.nlm.nih.gov/genome/?term=Hydra) and EST data bases10 (NCBI Hydra EST Database: https://www.ncbi.nlm.nih.gov/nucest), we now have a better understanding of the full spectrum of Hydra laminins. Based on current NCBI gene and EST data bases, Table 1 provides a list all identified Hydra laminins. Table 1 lists 6 Hydra laminin chains as analogs to vertebrate laminin chains. Some of these laminin chains were co-identified in both genomic studies and EST studies (laminin α1, laminin γ1). Other than Hydra laminin α1, and β1,13,23 the functions for Hydra laminin a2, a3, β1, b3, and γ1 chains cannot be known without further cellular and molecular analyses.
From available studies and data bases, it is clear that laminins emerged early during the formation of metazoans and in tandem with the appearance of other ECM components. This likely reflects the critical function of ECM in metazoans. ECM functions encompass complex processes involving not only the structural integrity of tissues, but the dynamic nature of ECM/cell interactions as related to cell physiological processes, cell differentiation, morphogenesis, and development in general.
The author wishes to thank those who have developed the genomic and EST data bases that have allowed more in depth study of the relationship of Hydra laminins to vertebrate laminins.
Author declares that there is no conflict of interest.

©2017 Sarras. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.