Journal of
eISSN: 2377-4312


Mini Review Volume 2 Issue 2
1Graduate Institute of Biotechnology, Chinese Culture University, Taiwan
2Department of Horticulture and Landscape Architecture, National Taiwan University, Taiwan
Correspondence: Yen-Chang Lin, 55. Hwa-Kang Rd. Yang-Ming-Shan. Taipei 11114, Taiwan, Tel 886 2 2861 0511, Fax 886 2 2861 8266
Received: July 17, 2014 | Published: April 9, 2015
Citation: Thi-Ngoc-Hieu B, Pung-Ling H, Shu-Yin W, et al. Associate micro-RNA expression profile with its target MRNA for new trend diagnosis and treatment in cancer. J Dairy Vet Anim Res. 2015;2(2):30-33. DOI: 10.15406/jdvar.2015.02.00026
MicroRNA has been focused recently in research, diagnosis and new-trend therapy in cancer. MicroRNA is known a short RNA (21-22 nucleotides) which plays important roles in cellular mechanisms can affected to cancer disease. Computation of numerous microRNA is not necessary; therefore, scientists concentrate on “seed-region” in microRNA structure and its targeted messenger RNA (mRNA). Remarkably experiments have been listed in this review show us how microRNA is working. These specific microRNAs are miR-155, miR-17~92 family, let-7 family, miR-15a~16.1 have been reported significant for some kinds of cancers. In the future, the evidences of this review hopefully can reduce duration of diagnosis and increase overall of cancer patients. MicroRNA and its targeted mRNA becomes a major tool in new recently approaching biomedical researches for pathology, human sciences.
Keywords: microrna, cancer, seed-region
NSCLS, non-small cell lung cancer; RISC, RNA induced silencing complex
Scientists have been overwhelming via processing regulated alterations of cellular mechanism activities chiefly from small component. Small fraction of RNAs, called microRNA (21~22 nucleotides) stimulates or inhibits differentiation, proliferation, apoptosis, stress responding and even tumor suppressor.1 Consequently, implicated modification interacts with DNA mutants, and alters protein levels of expression, as well as tumorigenesis. Many reports have been recorded about the functions of microRNA in evaluation and diagnosis of disease, especially in cancer.
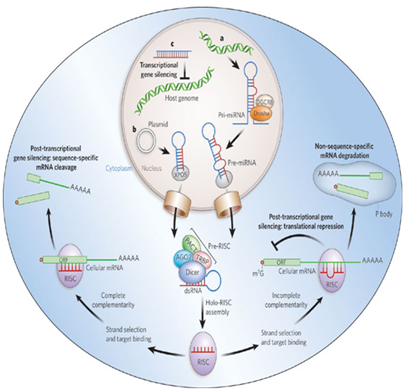
Twenty years ago the first two crucial microRNAs founded from the nematode Caenorhabditis elegans are Lin-4 and Let-7 which are the best structural characteristic of microRNA (Figure 1). Researchers have found hundreds of microRNAs in diverse species lately up to thousands. The most interesting fact is that many of the microRNAs in C.Elegans are abundant in humans, correlation that what is demonstrated in microRNA function in C.Elegans can be applied in human.2 A clearer figure of microRNA role plays an important affection in prognostic, diagnostic, and therapeutic for cancer diseases (Figure 2).
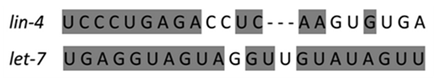
In recent years, estimated cancer mortalities due to cancer continue to rise (7.6million - 2008). GLOBOCAN evaluated in 27 sites of cancer in 184 countries and 12 world regions in 2008. The main contribution of death is lung, stomach and colorectal cancers in male while equivalently breast, colorectal and stomach in female.3 Oncologists, pathologists are now focusing on interrelated tumorigenesis factors researches. The fundamental circumstances associated with westernized lifestyles changing, infection-related cancer (H.Pylori in gastric cancer), reflected of microRNA performances. In this review, we have focused on the expression of microRNA levels.
Treatments for cancer nowadays are surgical, chemotherapy, radiotherapy comprehend accelerated overall ratio and non-sequelae in some patients. The prerequisite is how to return the diagnostic results quickly and to which therapy is corresponded to treat the patient. Prediction of microRNA roles in cellular mechanism activities and interaction with tumorigenesis which implement to diagnostic/prognostic have been reported in some researches.4 Evolutional discovery of microRNA develops new trend in biomedical cancer treatment to reduce duration of diagnostic, and then briefly conduct guideline relevant for each indication (Personal Health Care).
The progressing of mature microRNAs in bilateral symmetry involves in a chain of cleavage reactions conformity to primary microRNA (pri-miRNA) transcript.5 These pri-miRNAs are then clustered by complex of Endonuclease III enzyme called Drosha from hairpin form to double-stranded RNA-binding domain which is pre-miRNA.6 These pre-miRNAs are exported out of the nucleus to the cytoplasm via Exportin-5 than cleavage into single-stranded microRNA whereby Dicer.7 The mature microRNA composites into RNA induced silencing complex (called RISC) where the microRNA binding to target mRNA. Target recognition fragment on microRNA called “seed-region” 7 to 8 nucleotide d-long linked on 3 prime untranslated regions of mRNA that substitute differentiation durations and cellular mechanisms.8 Therefore, these alterations stimulate inhibited or blocked translation, accommodate protein expression related to cancer progression.
Affiliation between Laboratory of Immunogenomics, Zhoushan Hospital and CAS Key Laboratory of Genome Sciences and Information, Beijing Institute of Genomics had demonstrated that 2.24% up-regulated and 3.70% down-regulated target genes in total 34,694 investigated genes. The foremost figure for their experimental project which TargetScan and miRanda to predictively conjugate miRNA and mRNA between cancer tissues and controls. Tissues, contiguous tissues and sera in lung had collected from 35 non-small cell lung cancer (NSCLC) patients and 20 normal persons as control. Correspondence miRNAs and mRNAs expression in total RNA-extraction by RT-PCR shown that hsa-miR-96 represents an important role in NSCLC development as prediction.9 Subsequently, cellular function of hsa-miR-96 would be deepening research to elucidate correlation with targeted mRNA alterable expression, DNA methylation and histone modification so then reveal significant biomarkers for early diagnosis and comportable prognosis to increase survival rate in NSCLC patients.10
In addition, Sohila Zadran and his collaborate’s researching crucial tool miRNA and mRNA expression for diagnosis and prognosis in prostate, breast, lung, and ovarian cancer (Figure 3). Microarray and deep sequencing techniques had been used to detect mRNA and microRNA for this research. To interpret target, 8 healthy and 13 diseased patients for ovarian cancer, 37 healthy and 140 diseased patients for prostate cancer, 20 healthy and 20 diseased patients for breast cancer, 15 healthy and 15 diseased patients for breast cancer had been gathered to detect mRNA and miRNA signature (Figure 4). Surprisingly, in four different kind of cancers have equivalently remarkable results with mRNA down-regulated in diseased patients and mRNA up-regulated in healthy. On the other hand, significant miRNA for each disease has been revealed as correlation with mRNA expressions in ovarian cancer which is hsa-miR-135b; in prostate cancer which are hsa-miR-615, hsa-miR-3652, hsa-miR-153-1, hsa-miR-153-2, hsa-miR-1255a; in breast cancer which are hsa-miR-383, hsa-miR-1262; in lung cancer which are hsa-miR-1269, hsa-miR-577, hsa-miR-105-2.11 Consequently, it is probably many factors related to tumorigenesis can be affected by many miRNA as a balance network of cellular constituents. For abnormal condition, loss of one miRNA can increased another miRNAs to affect on targeted mRNA cause alterative in cellular mechanism. This research support that miRNA functions still is conspiratorial scenario should be sharpened resolve.12
Scientists combine modernized methodology to roll up mysterious microRNAs scenario. Computations and evaluations have found more than thousands of microRNA in all species. Biomedical knowledge prefer to know prerequisite accurate of specific microRNA roles and its target mRNA (Table 1). MicroRNA has been reported as a critical implement of alteration cellular mechanisms as activation of tumor processing, oncogenes, inhibition of developments and diseases, especially in cancer.
MicroRNA |
Affected cellular mechanisms |
References |
miR-155 |
Increase in several hematopoietic malignancies and tumors. |
Eis et al. (13) |
miR-17~92 |
Lung cancers |
Hayashita et al.(15) |
|
Develop a lymphoproliferative disorder affecting both B & T cells in autoimmunity |
Xiao et al. (19) |
miR-15a~16-1 |
Mantle cell lymphomas, prostate cancer |
Georges et al. (20) Bonci et al. (22) |
let-7 |
Induce cell cycle exit and terminal differentiation of cells |
Reinhart et al. (23) |
miR-10b |
Promotes cellular invasion and metastatic spread of transplanted tumors cause repression of protein HOXD10 |
Ma L et al. (26) |
miR-373 |
Promote cell migration in vitro |
Huang et al. (27) |
miR-200 |
Induce the EMT, deduce cellular migration and invasion |
Burk et al. (29) |
miR-126, miR-206, miR-335 |
Decrease expression resemble to poor metastasis-free in breast cancer patient. |
Tarvazoie et al. (31) |
Table 1 Affected cellular mechanisms
None.
Author declares that there is no conflict of interest.

©2015 Thi-Ngoc-Hieu, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.