Advances in
eISSN: 2572-8490


Objective: Melanoma, the most dangerous form of skin cancer, develops when unrepaired DNA damage to skin cells triggers mutations that lead the skin cells to multiply rapidly and form malignant tumours. The aim of this study was to evaluate the role of Micro RNA Let-7b in melanoma cell migration.
Materials and Methods: Melanoma specimens were obtained from 30 patients with skin melanoma. The human melanoma cell line A375 was used as an in vitro model for melanoma. In the present study, the wound healing assay and real-time PCR were carried out to assess the migration ability, and to determine mRNA transcription profiles, respectively.
Result: Results from this study showed that let-7b expression is able to significantly inhibit A375 melanoma cell migration. The wound healing assay demonstrated that A375 cell motility increases after let-7b inhibitor treatment.
Conclusion: Let-7b displayed a higher ability to inhibit melanoma cell migration and invasion, compared to other members of the Let-7 family.
Keywords: let-7b, melanoma, A375 cells
Melanoma is a type of cancer that develops from pigment-containing cells known as melanocytes.1 Melanomas typically occur in the skin but may rarely occur in the mouth and intestines.2 The legs and back are the most commonly involved sites in women and men, respectively.2 This type of cancer occasionally develops from a mole with worrying changes, including an increase in size and colour, irregular edges, itchiness or skin breakdown.3 Cancer is a multi-step process resulting from a gradual accumulation of genetic alterations caused by DNA damage.4 Thymine-thymine dimmers are the most type of DNA damage caused by ultraviolet A (UVA) light.3 More importantly, UVA produces reactive oxygen species (ROS) that induces DNA damage, including single-strand breaks, oxidized pyrimidine and oxidized purine 8-oxoguanine.2 It was shown that Micro RNAs (miRNAs) play an important role in cancer etiology and are frequently deleted in solid cancers including melanoma.5 miRNAs are small non-coding RNAs encoded in the human genome as miRNA genes, composing of either a single miRNA species (monocistronic) or several different miRNAs (polycistronic).5 MiR155, MiR146 and Let7 are the most miRNAs involved in melanoma.6 The Let-7 miRNA precursor was first identified in a study of developmental timing in C. elegans; however, it was later shown to be part of a much larger class of non-coding RNAs termed microRNAs.7 Importantly, Let-7 is a highly attractive potential therapeutic option with the ability to prevent tumor genesis and angiogenesis, typically in cancers.7 Let-7 has been demonstrated to be a direct regulator of RAS expression in human cells.8 All three RAS genes in human, including K-, N-, and H-ras, have the predicted let-7 binding sequences in their 3'UTRs.8 In samples from lung cancer patients, expression of RAS and let-7 showed a reciprocal pattern, in which cancerous cells have decreased let-7 and increased RAS; however, normal cells have increased let-7 and decreased RAS.2 Another oncogene, high mobility group A2 (HMGA2), has also been identified as a target of let-7.9 Interestingly, let-7 directly inhibits HMGA2 by binding to its 3'UTR. Removal of the let-7 binding site by 3'UTR deletion leads to HMGA2 over expression and tumor formation.9 In light of this, the purpose of this study was to investigate the Let-7b role in melanoma cell migration.
Patients and specimens
A total of 30 melanoma samples were collected from patients treated in the Sina Hospital, Tabriz University of Medical sciences, Tabriz, Iran. Tumors from primary sites were immediately snap-frozen and stored at −80°C.
Cell culture
The human melanoma cell line A375 was purchased from National Cell Bank of Iran (NCBI), Pasteur Institute of Iran, Tehran, Iran, and passaged in our laboratory for fewer than 6 months after resuscitation. The cells were cultured in RPMI 1640 medium (Cellgro, Manassas, VA, USA) supplemented with 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA) and 1% penicillin/streptomycin (Invitrogen, Carlsbad, CA, USA). The cell cultures were maintained in a humidified air chamber at 37°C with 5% CO2.
Migration and invasion assays
A375 human melanoma cells were transfected with let-7b mimic (60nM), let-7b inhibitor (90nM), and control oligos. Forty-eight hours following transfection, culture media were removed and the cells were serum-starved for 24 hours. Seventy-two hours after transfection, cells were trypsinized, suspended in serum-free medium, and then seeded into Trans well Permeable Support inserts with 8-μm micro-porous membranes (Corning, Corning, NY, USA) at a density of 2×105 cells per well in a 24-well plate. Culture medium containing 10% FBS was in the lower compartment of the plate. Six hours after culture, the non-migrating cells on the upper surface of the membrane were removed using a cotton swab. The cells that migrated to the lower surface were fixed and stained with the 3-step stain kit (Richard-Allan Scientific, Kalamazoo, MI, USA). The stained cells were photographed and counted with a haemocytometer under a microscope.
Wound healing assays
A375 human melanoma cells were seeded at a density of 1×106 cells per well into six-well plates and grown overnight. Cells were transfected with let-7b mimic (60 nM) and control oligos. After 48hours, the cells were serum starved for additional 24 hours. Afterwards, a sterile 200 μl pipette tip was applied to scratch the cells to form a wound. The cells were washed twice with PBS and then cultured in RPMI medium containing 10% FBS. Migration of the cells to the wound was observed and photographed using an inverted phase-contrast microscope equipped with digital camera system (Olympus). The healing rate was quantified by measuring the gap size after the culture. Eight different areas in each assay were chosen to evaluate the distance of the migrating cells to the origin of the wound. The healing rate of cells transfected with let-7b miRNA was normalized to that of control cells in a given experiment. The healing rate was used to represent cell migration.
Real-time PCR
Total RNA was extracted from the samples using TRIZOL reagent (Invitrogen, Carlsbad, CA, USA), and reverse-transcribed using a high capacity RNA-to-cDNA kit (Applied Bio systems, Foster City, CA, USA) according to the manufacturer's recommendation. The cDNA was quantified by RT-PCR using an ABI Prism 7900 sequence detection system (Applied Bio systems, Foster City, CA, USA). PCR amplification was carried out using SYBR Green PCR core reagents (Applied Bio systems, Foster City, CA, USA) according to the manufacturer’s instructions. PCR amplification of the housekeeping gene B-actin was carried out as a control for each sample.
Statistical analysis
All data were represented as mean ± SD. Statistical analysis was conducted using the Student’s test. A P value less than 0.05 was considered to be statistically significant.
let-7b levels in melanoma patients and its effect on A375 human melanoma cell migration
A decrease in let-7b levels was found in melanoma patients. let-7b was able to inhibit A375 human melanoma cell migration. To determine the clinical significance of the let-7 family in melanoma, the expression of all let-7 family members was also analyzed in a total 30 melanoma specimens using real-time RT-PCR. Results from real-time PCR summarized in Figure 1A indicate the lower expression levels of let-7b in melanoma cells compared to other members of the family. Moreover, it should be noted that B-actin was used as a control in this study, as shown in Figure 1A. The results strongly suggested that decreased expression of let-7 is an important step in melanoma development. To examine the effect of let-7b on A375 melanoma cell migration and invasion, the let-7b mimic or control were transfected into A375 melanoma cells. The migration and invasion abilities of transfected cells were examined 48 hours after transfection. Our results demonstrated that let-7b expression was able to significantly inhibit A375 melanoma cell migration (p<0.005; Figure 1B) and invasion (p<0.005; Figure 1C) in vitro.
The let-7b inhibitor was used to block endogenous let-7b expression. Results from the loss-of-function experiment demonstrated that the let-7b inhibitor significantly increases A375 melanoma cell migration (p<0.01; Figure 1D) and invasion (p<0.01; Figure 1E) in vitro.
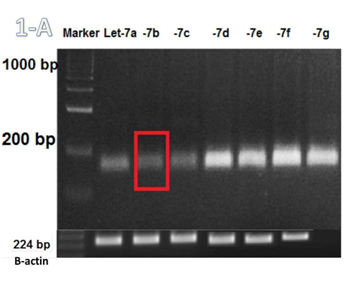
Figure 1A The PCR imaging of Let-7 family expression: As shown the let7-b expression is less than other family.
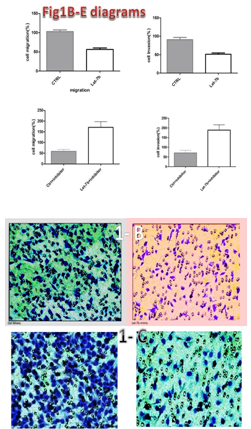
Figures 1B & 1C Cell migration and invasion percentages and results: let-7b expression was able to significantly inhibit A375 melanoma cell migration (p<0.005; Figure 1B) and invasion (p<0.005; Figure 1C).
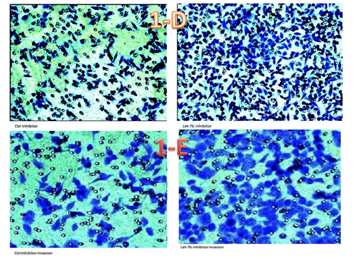
Figure 1D & 1E Cell migration and invasion percentages with using inhibitor: let-7b inhibitor significantly increases A375 melanoma cell migration (p<0.01; Figure 1D) and invasion (p<0.01; Figure 1E).
The effect of let-7b in melanoma cell motility
Cell motility is a factor that can impact cell migration in cancer. Results from the wound healing assay demonstrated that A375 cell motility increases after let-7b inhibitor treatment (Figure 2), emphasizing the inhibitory role of let-7b in melanoma cell motility.
Melanoma is the rarest form of skin cancer, accounting for only about 4% of skin cancer cases but for as many as 74% of all skin cancer deaths.10 Melanoma is more likely to invade nearby tissues and spread to other parts of the body than other types of skin cancer.11 MiRNAs let-7 is an important tumor suppressor gene in human. The let-7 family is a large family with thirteen members in human genome.7 Most studies demonstrated that let-7a is down-regulated in the tumor metastasis; however, few studies showed that there is a relation between let-7b and tumor growth.12 There is evidence that Let-7 can regulate various steps of the metastasis cascade in cancer cells.12 Our results reveal that let-7b could suppress melanoma cell migration and motility. Consistent with our results, a variety of studies showed the same function of let-7a in other cancers.13 The authors stated that only let-7e, among let-7 families, has no managing role in cancer metastasis. Results from the present study demonstrated that cell motility increases after let-7b inhibitor treatment. Taken together, the results from this investigation indicate that the Let-7 family, particularly Let-7b, could inhibit cell migration in malignant melanoma.
We would like to especially thank Prof Xu Huaxi for his advice in designing the study.
The author declares there is no conflict of interest.

© . This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.